이 번역 페이지는 최신 내용을 담고 있지 않습니다. 최신 내용을 영문으로 보려면 여기를 클릭하십시오.
plsregress
부분 최소제곱(PLS) 회귀
구문
설명
[는 다음 데이터도 반환합니다.XL,YL,XS,YS,BETA,PCTVAR,MSE,stats] = plsregress(X,Y,ncomp)
예측 변수 점수
XS. 예측 변수 점수는X에 있는 변수의 선형 결합인 PLS 성분입니다.응답 변수 점수
YS. 응답 변수 점수는 PLS 성분XS가 최대 공분산을 갖는 응답 변수의 선형 결합입니다.PLS 회귀 모델에 대한 계수 추정값으로 구성된 행렬
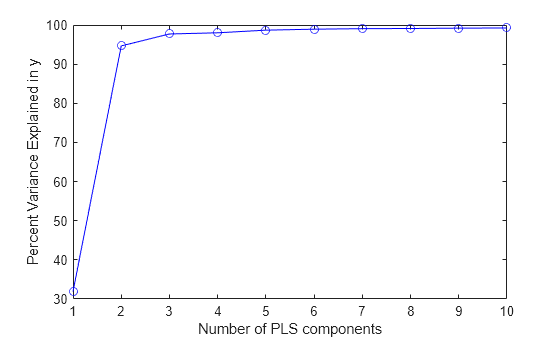
BETA회귀 모델로 설명되는 분산
PCTVAR의 백분율ncomp성분을 가진 PLS 모델에 대한 추정된 평균제곱오차MSEPLS 가중치, T2 통계량, 예측 변수 잔차와 응답 변수 잔차를 포함하는 구조체
stats.
예제
입력 인수
이름-값 인수
출력 인수
알고리즘
plsregress는 SIMPLS 알고리즘을 사용합니다[1]. 모델 피팅이 상수항(절편)을 포함하는 경우, 이 함수는 먼저 열 평균을 빼서 X와 Y를 중심화하고 중심화된 예측 변수와 응답 변수(각각 X0와 Y0)를 구합니다. 그러나 이 함수는 열을 다시 스케일링하지 않습니다. 표준화된 변수로 PLS 회귀를 수행하려면 zscore를 사용하여 X와 Y를 정규화합니다(X0 및 Y0의 열이 평균 0을 갖도록 중심화되고 표준편차 1을 갖도록 스케일링됨).
X와 Y를 중심화한 후, plsregress는 X0'*Y0에 대한 특이값 분해(SVD)를 계산합니다. 예측 변수 적재값과 응답 변수 적재값(각각 XL과 YL)은 예측 변수 점수 XS에 대해 X0과 Y0을 회귀 분석하여 얻은 계수입니다. XS*XL'과 XS*YL'을 각각 사용하여, 중심화된 데이터 X0 및 Y0을 복원할 수 있습니다.
plsregress는 처음에 YS를 YS = Y0*YL로 계산합니다. 하지만 일반적으로[1], plsregress는 XS'*YS가 하부 삼각 행렬이 되도록 XS의 이전 열을 기준으로 YS의 각 열을 직교화합니다.
모델 피팅이 상수항(절편)을 포함하지 않은 경우 X와 Y는 피팅 과정의 일부로 중심화되지 않습니다.
참고 문헌
[1] de Jong, Sijmen. “SIMPLS: An Alternative Approach to Partial Least Squares Regression.” Chemometrics and Intelligent Laboratory Systems 18, no. 3 (March 1993): 251–63. https://doi.org/10.1016/0169-7439(93)85002-X.
[2] Rosipal, Roman, and Nicole Kramer. "Overview and Recent Advances in Partial Least Squares." Subspace, Latent Structure and Feature Selection: Statistical and Optimization Perspectives Workshop (SLSFS 2005), Revised Selected Papers (Lecture Notes in Computer Science 3940). Berlin, Germany: Springer-Verlag, 2006, vol. 3940, pp. 34–51. https://doi.org/10.1007/11752790_2.
[3] Chong, Il-Gyo, and Chi-Hyuck Jun. “Performance of Some Variable Selection Methods When Multicollinearity Is Present.” Chemometrics and Intelligent Laboratory Systems 78, no. 1–2 (July 2005) 103–12. https://doi.org/10.1016/j.chemolab.2004.12.011.
확장 기능
버전 내역
R2008a에 개발됨