signalDatastore
Datastore for collection of signals
Description
Use a signalDatastore object to manage a collection of in-memory
data or signal files, where each individual file fits in memory, but the entire collection
does not necessarily fit.
Creation
Syntax
Description
sds = signalDatastore(data)data.
sds = signalDatastore(location)location. If location contains a mixture of
MAT files and CSV files, then sds contains MAT files.
sds = signalDatastore(___,Name,Value)
Input Arguments
In-memory input data, specified as vectors, matrices, timetables, or cell arrays.
Each element of data is a member that is output by the datastore
on each call to read.
Example: {randn(100,1); randn(120,3); randn(135,2);
randn(100,1)}
Files or folders to include in the datastore, specified as one of these values:
FileSetobject — Specifying the location as aFileSetobject leads to a faster construction time for datastores compared to specifying a path orDsFileSetobject. For more information, seematlab.io.datastore.FileSet.DsFileSetobject — For more information, seematlab.io.datastore.DsFileSet.File path — You can specify a single file path as a string scalar or character vector. You can specify multiple file paths as a string array or cell array of character vectors.
Files or folders can be local or remote:
Local files or folders — If the files are not in the current folder, then specify full or relative paths. Files within subfolders of a specified folder are not automatically included in the datastore. You can use the wildcard character (*) when specifying the local path. This character specifies that the datastore include all matching files or all files in the matching folders.
Remote files or folders — Specify full paths to remote files or folders as a uniform resource locator (URL) of the form
hdfs:///. Internet URLs must include the protocol typepath_to_file"http://"or"https://". For more information, see Work with Remote Data.
When you specify a folder, the datastore includes only files with supported file
formats and ignores files with any other format. To specify a custom list of file extensions to
include in your datastore, see the FileExtensions name-value argument.
Example: 'whale.mat'
Example: '../dir/data/signal.mat'
Data Types: char | string | cell
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Before R2021a, use commas to separate each name and value, and enclose
Name in quotes.
Example: sds =
signalDatastore('C:\dir\signaldata','FileExtensions','.csv')
Subfolder inclusion flag, specified as true or
false. Specify true to include all files and
subfolders within each folder or false to include only the files
within each folder.
Example: 'IncludeSubfolders',true
Data Types: logical | double
Signal file extensions, specified as a string scalar, string array, character vector, or cell array of character vectors.
If no read function is specified, 'FileExtensions' can only
be set to .mat to read MAT files, or to .csv
to read CSV files. If 'FileExtensions' is omitted, it defaults
to .mat if there are MAT files in the specified location,
otherwise 'FileExtensions' defaults to .csv
if there are CSV files in the specified location.
If the specified location contains both MAT files and CSV files,
signalDatastore defaults to reading the MAT files. If neither MAT
files nor CSV files are present, signalDatastore errors out with the
default read function. Specify a custom
read using ReadFcn function to read files of any other type.
When you do not specify a file extension, the signalDatastore needs
to parse the files to decide the default extension to read. Specify an extension to
avoid the parsing time.
Example: 'FileExtensions','.csv'
Data Types: string | char | cell
In addition to these name-value arguments, you also can specify any of the properties
on this page as name-value arguments, except for the Files
property.
Properties
In-Memory Data
Member names, specified as a cell array. The length of the member names for the
input data should equal the length of the data cell array. This
property applies only when the datastore contains in-memory data.
Signal member data, specified as a string scalar or a string array. The length of
the member names for the input data should equal the length of the
data cell array. This property applies only when the datastore
contains in-memory data.
File Data
Files included in the datastore, specified as a cell array of strings or character
vectors. Each character vector in the cell array represents the full path to a file.
The location argument in the signalDatastore
defines Files when the datastore is created. This property
applies only when the datastore contains file data.
Data Types: string | char | cell
Function that reads data, specified as a function handle. The function must take a
file name as input, and then it outputs the corresponding data. For example, if
customreader is the specified function to read the data, then it
must have one of these
templates:
function data = customreader(filename) ... end
function [data,info] = customreader(filename) ... end
data variable. The
info variable must be a user-defined structure containing
user-defined information from the file. If you need extra arguments, you can include
them after the filename argument. signalDatastore
appends to the info structure a field containing the name of the
file.Example: @customreader
Data Types: function_handle
Alternate file system root paths, specified as the name-value argument consisting of
"AlternateFileSystemRoots" and a string vector or a cell array. Use
"AlternateFileSystemRoots" when you create a datastore on a local
machine, but need to access and process the data on another machine (possibly of a different
operating system). Also, when processing data using the Parallel Computing Toolbox™ and the MATLAB®
Parallel Server™, and the data is stored on your local machines with a copy of the data available
on different platform cloud or cluster machines, you must use
"AlternateFileSystemRoots" to associate the root paths.
To associate a set of root paths that are equivalent to one another, specify
"AlternateFileSystemRoots"as a string vector. For example,["Z:\datasets","/mynetwork/datasets"]
To associate multiple sets of root paths that are equivalent for the datastore, specify
"AlternateFileSystemRoots"as a cell array containing multiple rows where each row represents a set of equivalent root paths. Specify each row in the cell array as either a string vector or a cell array of character vectors. For example:Specify
"AlternateFileSystemRoots"as a cell array of string vectors.{["Z:\datasets", "/mynetwork/datasets"];... ["Y:\datasets", "/mynetwork2/datasets","S:\datasets"]}Alternatively, specify
"AlternateFileSystemRoots"as a cell array of cell array of character vectors.{{'Z:\datasets','/mynetwork/datasets'};... {'Y:\datasets', '/mynetwork2/datasets','S:\datasets'}}
The value of "AlternateFileSystemRoots" must satisfy these conditions:
Contains one or more rows, where each row specifies a set of equivalent root paths.
Each row specifies multiple root paths and each root path must contain at least two characters.
Root paths are unique and are not subfolders of one another.
Contains at least one root path entry that points to the location of the files.
For more information, see Set Up Datastore for Processing on Different Machines or Clusters.
Example: ["Z:\datasets","/mynetwork/datasets"]
Data Types: string | cell
Names of variables in signal files, specified as a string scalar or vector of unique names. Use this property when your files contain more than one variable and you want to specify the names of the variables that hold the signal data you want to read.
When the property value is a string scalar,
signalDatastorereturns data contained in the specified variable.When the property value is a string vector,
signalDatastorereturns a cell array with the data contained in the specified variables. In this case, you can use theReadOutputOrientationproperty to specify the orientation of the output cell array as a column or a row.
If this property is not specified, signalDatastore reads
the first variable in the variable list of each file.
Note
To determine the name of the first variable in a file,
signalDatastore follows these steps:
For MAT files:
s = load(fileName); varNames = fieldnames(s); firstVar = s.(varNames{1});For CSV files:
opts = detectImportOptions(fileName,'PreserveVariableNames',true); varNames = opts.VariableNames; firstVar = string(varNames{1});
This property applies only when the datastore contains file data and the default read function is used.
Output signal data cell array orientation, specified as
'column' or 'row'. This property specifies how
to orient the output signal data cell array after a call to the read
function when SignalVariableNames contains more than one signal
name. ReadOutputOrientation has no effect when
SignalVariableNames contains only one element and does not
apply if SignalVariableNames has not been specified.
This property applies only when the datastore contains file data and the default read function is used.
Example: Output Cell Array Orientation
In the Read Multiple Variables from Files in Signal Datastore example,
data has the default output orientation and is a 2-by-1 column
array:
{1×4941 double}
{1×4941 double}
ReadOutputOrientation as 'row',
then data is a 1-by-2 row
array: {1×4941 double} {1×4941 double}Name of the variable holding the sample rate, specified as a string scalar. This property applies only when the datastore contains file data.
Name of the variable holding the sample time value, specified as a string scalar. This property applies only when the datastore contains file data.
Name of the variable holding the time values vector, specified as a string scalar. This property applies only when the datastore contains file data.
Note
'SampleRateVariableName',
'SampleTimeVariableName', and
'TimeValuesVariableName' are mutually exclusive. Use these
properties when your files contain a variable that holds the time information of the
signal data. If not specified, signalDatastore assumes that signal data has
no time information. These properties are not valid if a custom read
function is specified.
In-Memory and File Data
Sample rate values, specified as a positive real scalar or vector.
Set the value of
SampleRateto a scalar to specify the same sample rate for all signals in thesignalDatastore.Set the value of
SampleRateto a vector to specify a different sample rate for each signal in thesignalDatastore.
The number of elements in the vector must equal the number of elements
in the signalDatastore.
Sample time values, specified as a positive scalar, a vector, a duration scalar, or a duration vector.
Set the value of
SampleTimeto a scalar to specify the same sample time for all signals in thesignalDatastore.Set the value of
SampleTimeto a vector to specify a different sample time for each signal in thesignalDatastore.
The number of elements in the vector must equal the number of elements
in the signalDatastore.
Time values, specified as a vector, a duration vector, a matrix, or a cell array.
Set
TimeValuesto a numeric ordurationvector to specify the same time values for all signals in thesignalDatastore. The vector must have the same length as all the signals in the set.Set
TimeValuesto a numeric ordurationmatrix or cell array to specify that each signal of thesignalDatastorehas signals with the same time values, but the time values differ from signal to signal.If
TimeValuesis a matrix, then the number of columns equal the number of members of thesignalDatastore. All signals in the datastore must have a length equal to the number of rows of the matrix.If
TimeValuesis a cell array, then the number of vectors equal the number of members of thesignalDatastore. All signals in a member must have a length equal to the number of elements of the corresponding vector in the cell array.
Maximum number of signal files returned by read, specified as
a positive real scalar. If you set the ReadSize property to n, such that
n > 1, each time you call the read
function, the function reads:
The first variable of the first n files, if
sdscontains file data.The first n members, if
sdscontains in-memory data.
The output of read is a cell array of signal data
when ReadSize > 1.
Since R2024b
Hardware resource for output data, specified as one of these:
"cpu"— Return read data on the CPU."gpu"— Return read numeric data on the GPU asgpuArrayobjects.
When you read each member of the signalDatastore object with the
OutputEnvironment property set to "gpu", the
read
function attempts moving the data from each member element into the GPU.
If the member element is numeric,
readmoves its data into the GPU using agpuArrayobject.If the member element is not numeric,
readkeeps its data in the CPU.
Using a GPU requires a Parallel Computing Toolbox license and a supported GPU device. For information on supported devices, see GPU Computing Requirements (Parallel Computing Toolbox).
Data Types: char | string
Since R2024b
Data type of read output, specified as one of these:
"same"— Do not cast data and return."double"— Cast read data to double precision."single"— Cast read data to single precision.String array or cell array of character vectors — Cast read data from each member element and return with the specified new data type.
You must create the
signalDatastoreobject from file data to specifyOutputDataTypeas a string array or cell array of character vectors.Specify
OutputDataTypeas an array where each element is one of these:"same","double", or"single".The number of elements in the array specified in
OutputDataTypemust be 1, or must correspond with the number of variables specified inSignalVariableNames.
When you read each member of the signalDatastore object with the
OutputDataType property set to "single" or
"double", the read
function attempts casting all member elements of the object to single-precision or
double-precision numbers.
If all the member elements support conversion to the specified new data type,
readconverts and returns each element with the specified new data type.If any member element does not support conversion to the specified new data type,
readerrors out.
When you specify the OutputDataType property with a string
array or cell array of character vectors, the read function
attempts casting the data from each variable specified in
SignalVariableNames depending on the number of elements in
OutputDataType.
If
OutputDataTypehas one element, thereadfunction casts all the variables specified inSignalVariableNamesto the data type specified inOutputDataTypeif all the variables support conversion to the specified new data type.If
OutputDataTypehas as many elements as variables specified inSignalVariableNames, thereadfunction casts the ith variable inSignalVariableNamesto the ith data type specified inOutputDataTypeif the ith variable supports conversion to the specified new data type.If any variable does not support conversion to the specified new data type,
readerrors out.
For more information about determining data casting support, see cast.
Data Types: char | string
Object Functions
read | Read next consecutive signal observation |
readall | Read all signals from datastore |
writeall | Write datastore to files |
preview | Read first signal observation from datastore for preview |
shuffle | Shuffle signals in signal datastore |
subset | Create datastore with subset of signals |
partition | Partition signal datastore and return partitioned portion |
numpartitions | Return estimate for reasonable number of partitions for parallel processing |
reset | Reset datastore to initial state |
progress | Determine how much data has been read |
hasdata | Determine if data is available to read |
transform | Transform datastore |
combine | Combine data from multiple datastores |
isPartitionable | Determine whether datastore is partitionable |
isShuffleable | Determine whether datastore is shuffleable |
Note
isPartitionable
and isShuffleable
return true by default for signalDatastore. You can test
if the output of combine and
transform are
partitionable or shuffleable using the two functions.
Examples
Create a signal datastore to iterate through the elements of an in-memory cell array of signal data. The data consists of a sinusoidally modulated linear chirp, a concave quadratic chirp, and a voltage controlled oscillator. The signals are sampled at 3000 Hz.
fs = 3000;
t = 0:1/fs:3-1/fs;
data = {chirp(t,300,t(end),800).*exp(2j*pi*10*cos(2*pi*2*t)); ...
2*chirp(t,200,t(end),1000,'quadratic',[],'concave'); ...
vco(sin(2*pi*t),[0.1 0.4]*fs,fs)};
sds = signalDatastore(data,'SampleRate',fs);While the datastore has data, read each observation from the signal datastore and plot the short-time Fourier transform.
plotID = 1; while hasdata(sds) [dataOut,info] = read(sds); subplot(3,1,plotID) stft(dataOut,info.SampleRate) plotID = plotID + 1; end
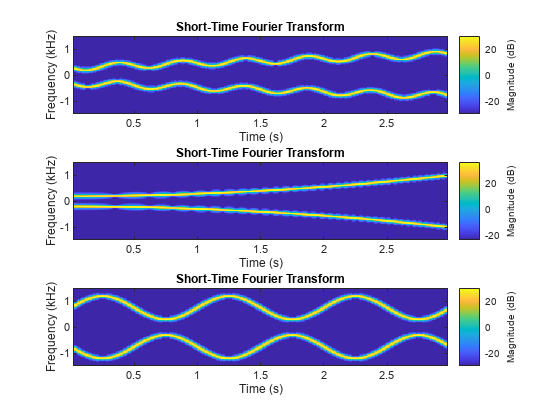
The folder dataset contains signal samples included with Signal Processing Toolbox™. Create a signal datastore that points to the folder and set the name of the sample rate variable.
folder = "dataset"; sds = signalDatastore(folder,SampleRateVariableName="fs");
Read the first file in the datastore and plot the spectrogram.
[data,info] = read(sds);
pspectrum(data,info.SampleRate,"spectrogram")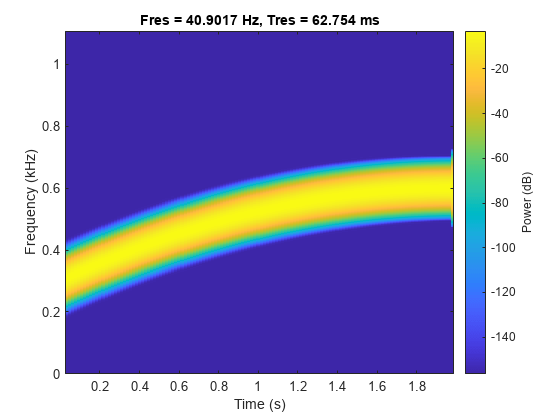
Specify the folder that contains signal samples included with Signal Processing Toolbox™. The signals are stored in .csv, .dat, and .mat files.
folder = "healthdata";Create a signal datastore that points to the .csv file in the specified folder. Plot the short-time Fourier transform of the signal.
sds = signalDatastore(folder, ... FileExtensions=".csv",SignalVariableNames=["tx" "x"]); data = read(sds); stft(data{2})
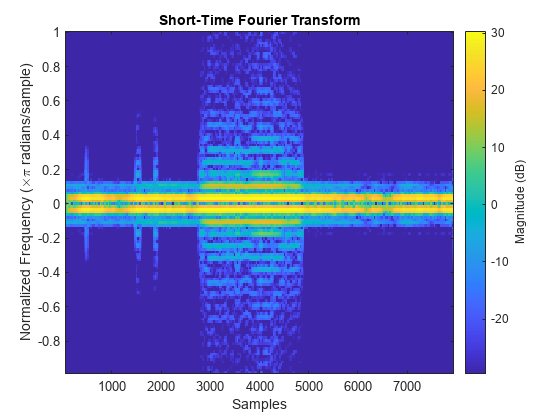
Specify the names of four example files included with Signal Processing Toolbox™.
files = ["INR.mat","relatedsig.mat","spots_num.mat","voice.mat"];
Create a signalDatastore object containing the specified files and set the ReadSize property to 2 to read data from two files at a time. Each read returns a cell array where the first cell contains the first variable of the first file read, and the second cell contains the first variable from the second file. While the datastore has data, display the names of the variables read in each read.
sds = signalDatastore(files,ReadSize=2); while hasdata(sds) [data,info] = read(sds); fprintf("Variable Name:\t%s\n",info.SignalVariableNames) end
Variable Name: Date Variable Name: s1 Variable Name: year Variable Name: fs
Create a signal datastore that contains three signals included with Signal Processing Toolbox™.
The
strong.matfile contains three variables:her,himandfs.The
slogan.matfile contains three variables:hotword,phraseandfs.The
Ring.matfile contains two variables:yandFs.
Each file contains multiple variables of different names. The scalar in each file represents a sample rate. Define a custom read function that reads all the variables in the file as a structure and returns the variable in dataOut and information about the variables in infoOut. The SampleRate field of infoOut contains the scalar contained in each file, and dataOut contains the variables read from each file.
function [dataOut,infoOut] = MyCustomRead(filename) fText = importdata(filename); value = struct2cell(fText); dataOut = {}; for i = 1:length(value) if isscalar(value{i}) == 1 infoOut.SampleRate = value{i}; else dataOut{end+1} = value{i}; end end end
files = ["strong.mat","slogan.mat","Ring.mat"]; sds = signalDatastore(files,ReadFcn=@MyCustomRead);
While the datastore has unread files, read from the datastore and compute the short-time Fourier transforms of the signals.
while hasdata(sds) [data,infoOut] = read(sds); fs = infoOut.SampleRate; figure for i = 1:length(data) if length(data)>1 subplot(2,1,i) end stft(data{i},fs) end end
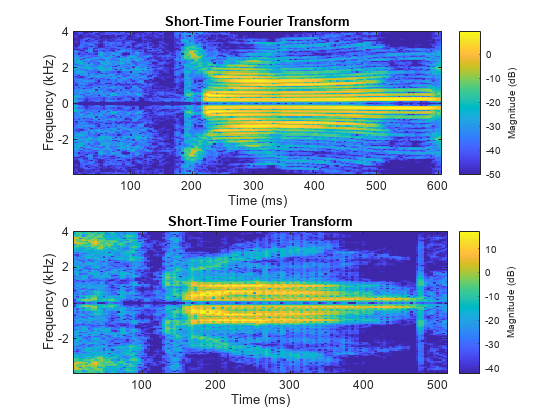

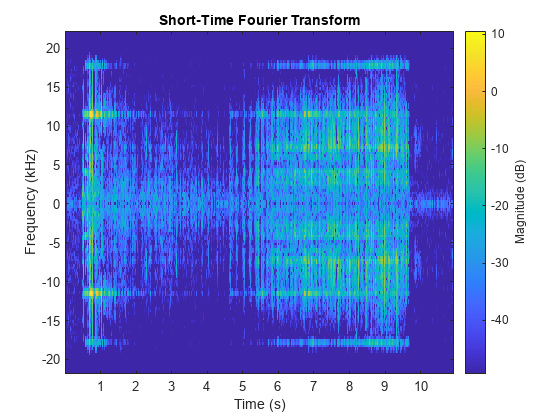
The dataset folder contains example files included with Signal Processing Toolbox™. Each file contains two signals and a random sample rate fs ranging from 3000 to 4000 Hz.
The first signal,
x1, is a convex quadratic chirp.The second signal,
x2, is a chirp with sinusoidally varying frequency content.
folder = "dataset";Create a signal datastore that points to the specified folder, set the names of the signal variables and sample rate, and specify the output data type as single precision. While the datastore has data, read each observation and visualize the spectrogram of each signal.
sds = signalDatastore(folder,SignalVariableNames=["x1";"x2"], ... SampleRateVariableName="fs",OutputDataType="single"); tiledlayout flow while hasdata(sds) [data,info] = read(sds); nexttile pspectrum(data{1},info.SampleRate,"spectrogram",TwoSided=true) nexttile pspectrum(data{2},info.SampleRate,"spectrogram",TwoSided=true) end

Extended Capabilities
The signalDatastore function
supports GPU array input with these usage notes and limitations:
This object can generate GPU arrays, but does not run on a GPU. A
signalDatastoreobject can return data on the GPU in agpuArrayobject if you set the OutputEnvironment property to"gpu".
For more information, see Run MATLAB Functions on a GPU (Parallel Computing Toolbox).
Version History
Introduced in R2020aWhen you use read or readall, you can now
select the precision of the signalDatastore output data and the hardware
that you want to use to return the output data. You must have Parallel Computing Toolbox to use gpuArray objects.
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
웹사이트 선택
번역된 콘텐츠를 보고 지역별 이벤트와 혜택을 살펴보려면 웹사이트를 선택하십시오. 현재 계신 지역에 따라 다음 웹사이트를 권장합니다:
또한 다음 목록에서 웹사이트를 선택하실 수도 있습니다.
사이트 성능 최적화 방법
최고의 사이트 성능을 위해 중국 사이트(중국어 또는 영어)를 선택하십시오. 현재 계신 지역에서는 다른 국가의 MathWorks 사이트 방문이 최적화되지 않았습니다.
미주
- América Latina (Español)
- Canada (English)
- United States (English)
유럽
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)