bwskel
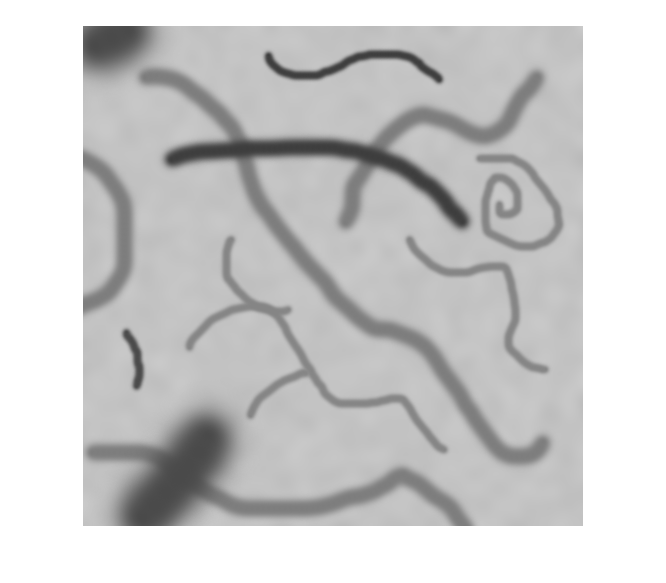
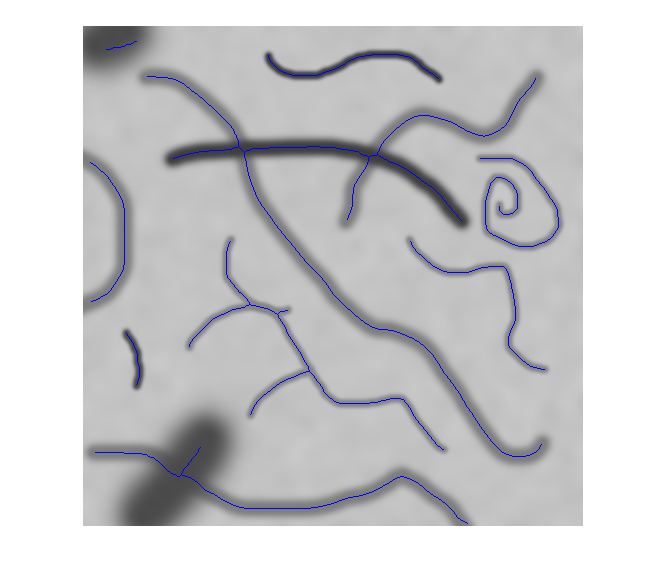
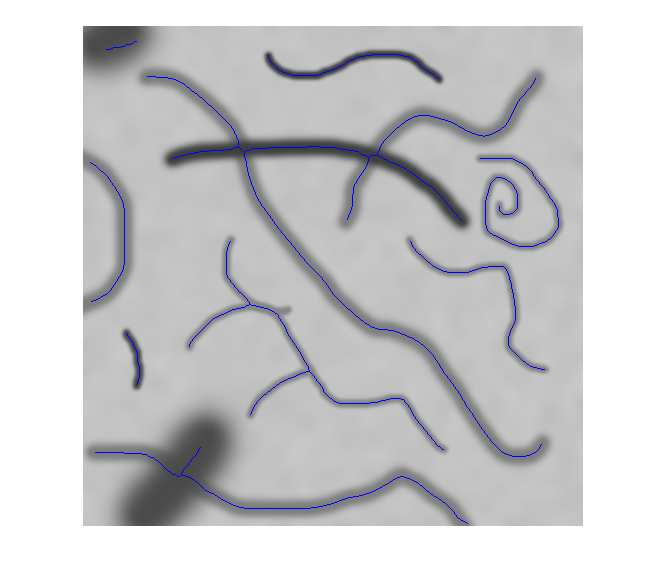
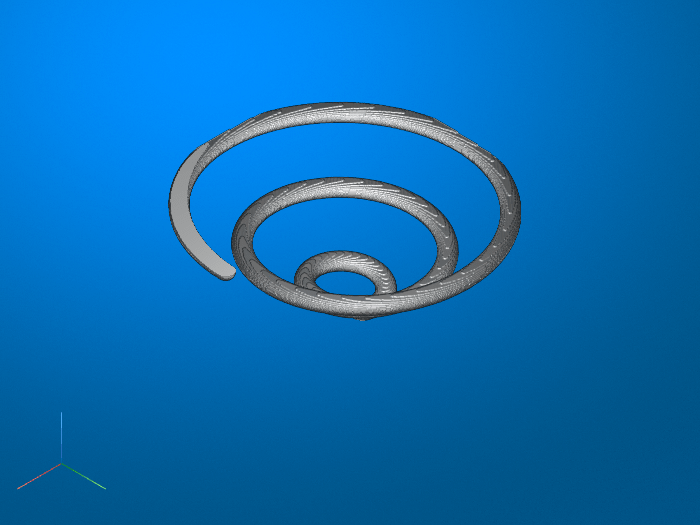
모든 객체를 2차원 이진 영상 또는 3차원 이진 볼륨의 선으로 축소
설명
예제
입력 인수
출력 인수
팁
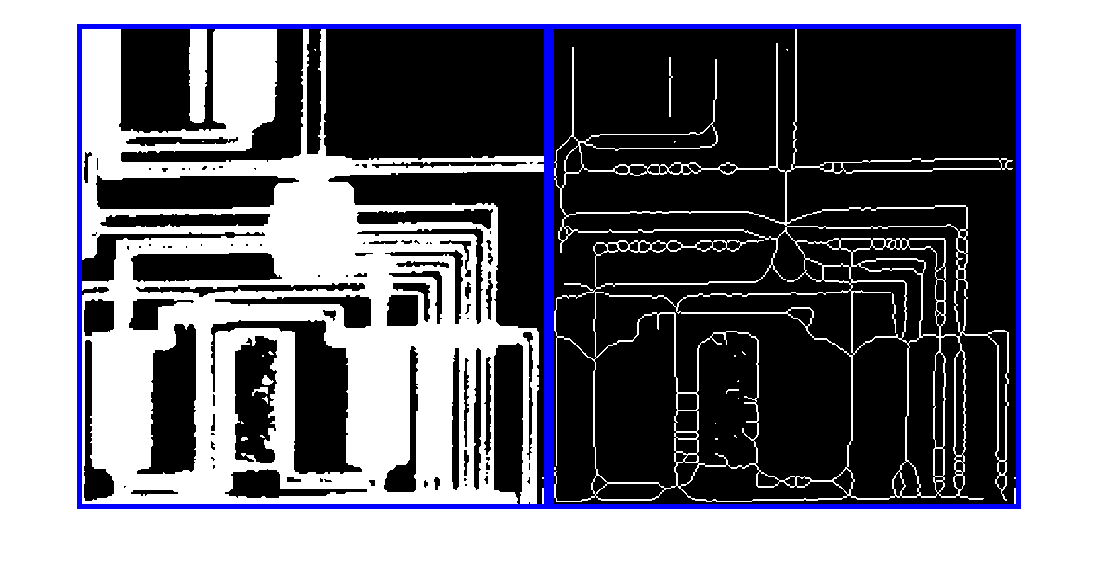
bwskel과bwmorph모두 2차원 영상을 골격화할 수 있지만, 두 함수는 서로 다른 알고리즘을 사용하므로 다른 결과를 반환할 수 있습니다.bwskel함수는 2차원 영상에 4-연결성을 사용하지만,bwmorph는 8-연결성을 사용합니다.bwmorph함수가 더 정확한 골격을 만드는 경우가 많습니다.bwskel은 이진 영상의 전경 객체가 흰색이라고 가정합니다(논리값true). 배경이 흰색이고 객체가 검은색인 영상이라면, 이 영상의 보수에 해당하는 색 반전 영상을bwskel에 대한 입력으로 사용하십시오. 색 반전은imcomplement함수를 사용하여 계산할 수 있습니다.
알고리즘
bwskel함수는 중앙 축 변환을 사용합니다.
참고 문헌
[1] Ta-Chih Lee, Rangasami L. Kashyap and Chong-Nam Chu. Building skeleton models via 3-D medial surface/axis thinning algorithms. Computer Vision, Graphics, and Image Processing, 56(6):462-478, 1994.
[2] Kerschnitzki, M, Kollmannsberger, P, Burghammer, M. et al. Architecture of the osteocyte network correlates with bone material quality. Journal of Bone and Mineral Research, 28(8):1837-1845, 2013.