Using SimBiology for Mechanism-Based PK/PD Modeling in Preclinical & Discovery
Drug developers are moving towards mechanism-based drug discovery, using mechanistic or semi-mechanistic models of drug action and efficiency to extend traditional PK modeling techniques. These mechanism-based models are more suitable for prediction and extrapolation than pure empirical approaches. In this webinar, you learn how to use SimBiology and MATLAB to implement mechanism-based PK/PD modeling workflows.
Using a modeling and simulation case-study from literature, we demonstrate:
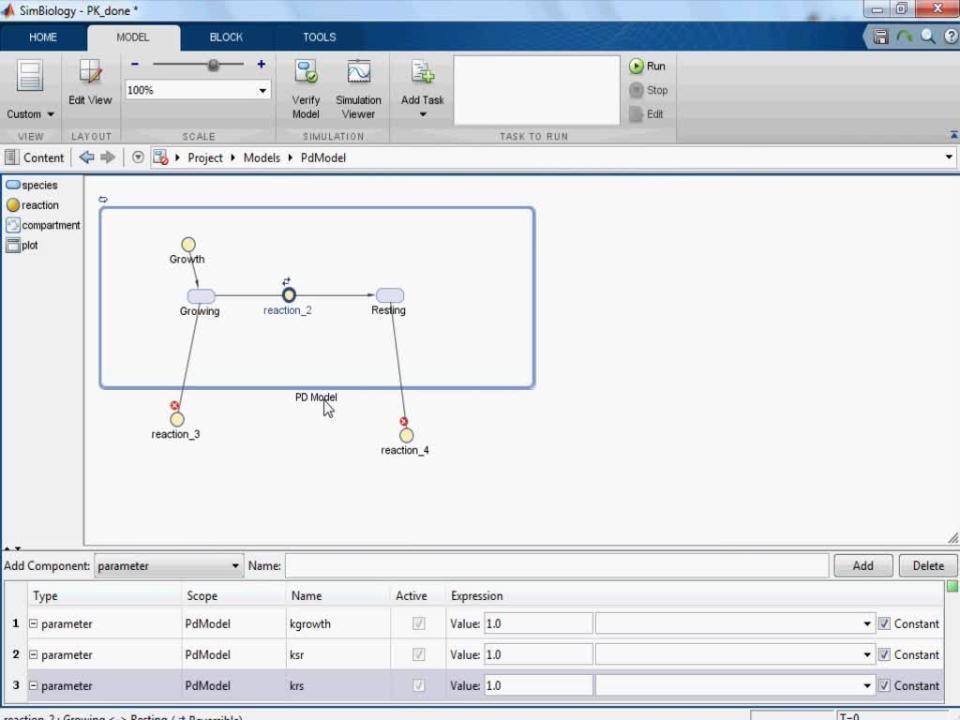
• Building mechanism-based PK/PD models using an interactive block-diagram editor
• Estimating parameters by fitting experimental data
• Simulating different dosing strategies
• Exploring system dynamics using parameter sweeps & sensitivity analysis
• Automating and customizing analyses using MATLAB
We also highlight new features in SimBiology 2012a including:
• Simulation Viewer – a interactive visualization tool for model exploration
• Weighted fitting
• Simultaneously fitting data from multiple dose levels
Recorded: 26 Mar 2012