Expression Analysis
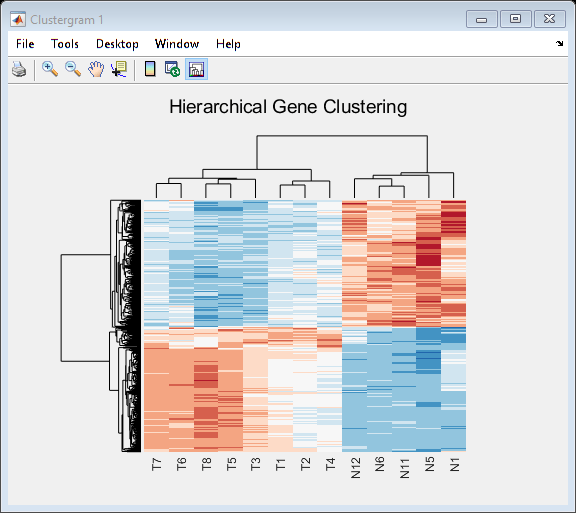
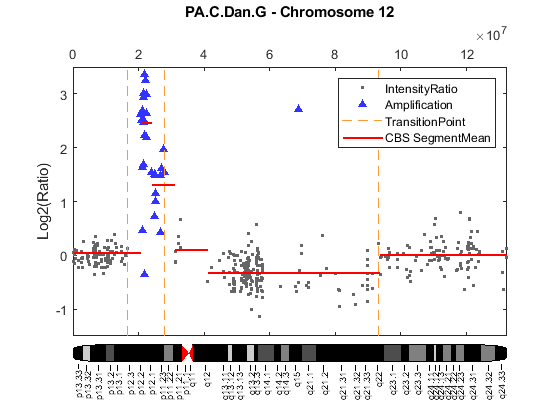
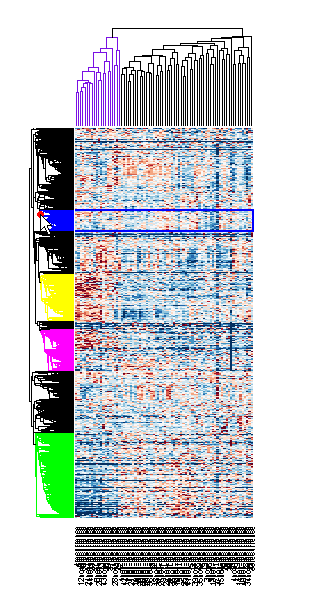
Evaluate gene expression data and identify differentially expressed genes using hypothesis tests. Use various machine learning functions to train classifier models, perform principal component analysis, rank features, and impute missing data. Visualize microarray data using intensity versus ratio (IR) plots, box plots, or log-log plots.
Functions
Objects
DataMatrix | Data structure encapsulating data and metadata from microarray experiment |
bioma.ExpressionSet | Contain data from microarray gene expression experiment |
bioma.data.ExptData | Contain data values from microarray experiment |
bioma.data.MetaData | Metadata from microarray experiment |
bioma.data.MIAME | Contain experiment information from microarray gene expression experiment |
NegativeBinomialTest | Unpaired hypothesis test result |
HeatMap | Object containing matrix and heatmap display properties |
clustergram | Object containing hierarchical clustering analysis data |
Topics
- Managing Gene Expression Data in Objects
Overview of objects for Microarray Gene Expression Data
- Representing Expression Data Values in DataMatrix Objects
Construct DataMatrix objects, get and set properties, and access data.
- Representing Expression Data Values in ExptData Objects
Construct ExptData objects, use properties and methods, and access data.
- Representing Sample and Feature Metadata in MetaData Objects
Construct MetaData objects, use properties and methods, and access data.
- Representing Experiment Information in a MIAME Object
Construct MIAME objects, use properties and methods, and access data.
- Representing All Data in an ExpressionSet Object
Construct ExpressionSet objects, use properties and methods, and access data.
- Microarray Data Analysis Tools
The MATLAB® environment is widely used for microarray data analysis, including reading, filtering, normalizing, and visualizing microarray data.
- Statistical Learning and Visualization
You can classify and identify features in data sets, set up cross-validation experiments, and compare different classification methods.