Label 2-D Ultrasound Series Using Medical Image Labeler
This example shows how to label 2-D image data using the Medical Image Labeler app. The app provides manual, semi-automated, and automated tools for labeling 2-D medical image data. This example labels the left ventricle in an echocardiogram ultrasound image series.
Open the Medical Image Labeler
Open the Medical Image Labeler app from the Apps tab on the MATLAB® Toolstrip, under Image Processing and Computer Vision. You can also load the app by using the medicalImageLabeler command.

Create New Image Labeling Session
To start a new 2-D labeling session, on the app toolstrip, click New Session and select New Image Session (2-D). In the Create a new session folder dialog box, specify a location in which to save the new session folder by entering a path or selecting Browse and navigating to your desired location. In the New Session Folder dialog box, specify a name for the folder for this labeling session. Then, select Create Session.
Load Image Data into Medical Image Labeler
To load an image into the Medical Image Labeler app, on the app toolstrip, click Import. Then, under Data, select From File. Browse to the location of heartUltrasoundSequenceVideo.dcm in the same directory as this example file. For an image session, the imported data file must be a single DICOM file, or a file in a format listed in the imformats file format registry, that contains a 2-D image or a series of 2-D images related by time. You can also import data as a medicalImage object from the workspace.
The name of the imported image is visible in the Data Browser pane. You can import multiple 2-D image files into an image session. All files imported into a single app session must label the same regions of interest, such as tumor or ventricle, and are exported together as one groundTruthMedical object.
Explore the Image Series
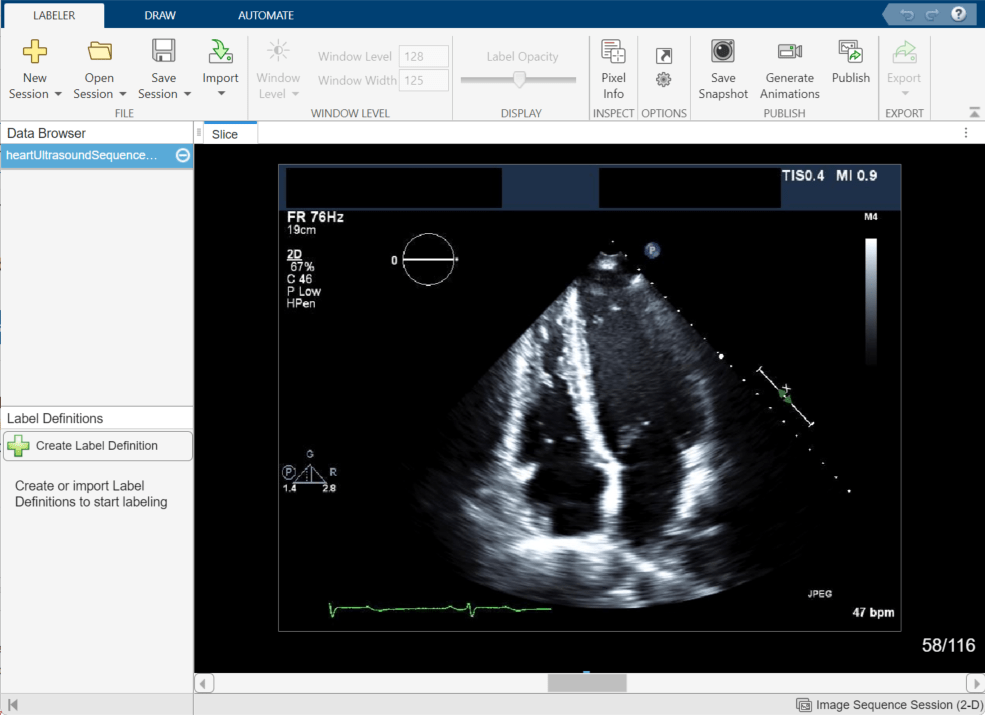
The Medical Image Labeler app displays the imported image in the Slice pane. By default, the app displays the middle frame in the ultrasound series. You can change the displayed frame by using the scroll bar at the bottom of the Slice pane, or you can click the pane and then press the left and right arrow keys. The app displays the current frame number out of the total number of frames, such as 58/116. You can zoom in on the current frame using the mouse scroll wheel or the zoom controls that appear when you pause on the Slice pane.
Explore the echocardiogram image series to identify the left ventricle. The annotated image shows the approximate outline of the ventricle to label. Note that all labels in this example have been created for illustrative purposes only and have not been verified by a clinical professional.

Create Label Definitions
A label definition specifies the name, color, and numeric index of a label. In the Label Definitions pane, select Create Label Definition to create a label with the default name Label1. To change the name of the label, click the label and enter a new name. The label name must be a valid MATLAB variable name with no spaces. For this example, specify the name of the label as LeftVentricle. To change the default color associated with the label, click the colored square in the label identifier and select a color from the Color dialog box.

Use Drawing Tools to Label Regions in Image
To assign pixels to the LeftVentricle label, click the LeftVentricle label in the Label Definitions pane. You can use the tools on the Draw tab in the app toolstrip to define the region. You can choose from the Freehand, Assisted Freehand, Polygon, Paint Brush, and Trace Boundary tools. Label frame 1. When you add the label, the app adds a bar above the slider, using the color associated with the label, to indicate which frames you have labeled.

In addition to the drawing tools, you can add or refine label regions using the tools in the Automate tab of the app toolstrip. The app provides automation algorithms including Active Contours, Adaptive Threshold, Dilate, and Erode. To apply an algorithm, click Algorithm Parameters to adjust settings, if applicable, and click Run. You can also specify a custom range of frames to process by specifying a Start frame and an End frame.
Use Interpolation to Speed Up Labeling
You can move through the image series and draw labels frame-by-frame, but the Medical Image Labeler app also provides interpolation tools in the Draw tab that can help you label an object between frames. Interpolation is most suitable between frames where the region of interest has a similar shape and size. In an echocardiogram, the ventricle experiences cycles of contraction, during which the ventricle rapidly changes in shape, and relaxation, during which the ventricle is relatively still. Therefore, you can most effectively use interpolation between the start and end of a relaxation period, such as between frames 20 and 38.
To use interpolation, you must first manually label a region in two frames. Label the ventricle in frame 20 and frame 38. If the Auto Interpolate button is not active, make sure the labeled region is selected by clicking Select Drawn Region in the app toolstrip and selecting the labeled region.
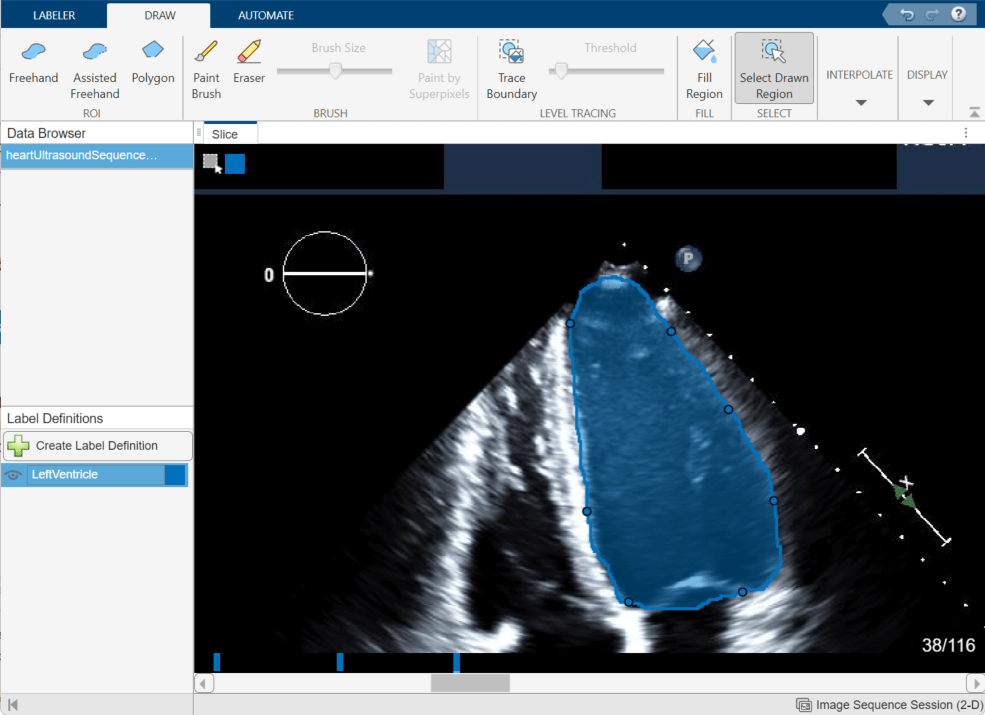
Click Auto Interpolate. The app automatically labels the ventricle in the intermediate frames. The app adds bars above the slider to indicate all the frames that have labeled pixels, which now appears as a solid bar from frame 20 to frame 38.
Alternatively, after labeling an ROI on two frames, you can click Manually Interpolate. With this option, the app opens the Manually Interpolate dialog box. Select the two regions between which you want to interpolate, Region One and Region Two. To select the first region, use the slider at the bottom of the dialog box to navigate to frame 20, and then click inside the labeled ventricle. To select the second region, click Region Two, navigate to frame 38, and click inside the labeled ventricle. After selecting both regions, click Run to interpolate the label in the intermediate frames.
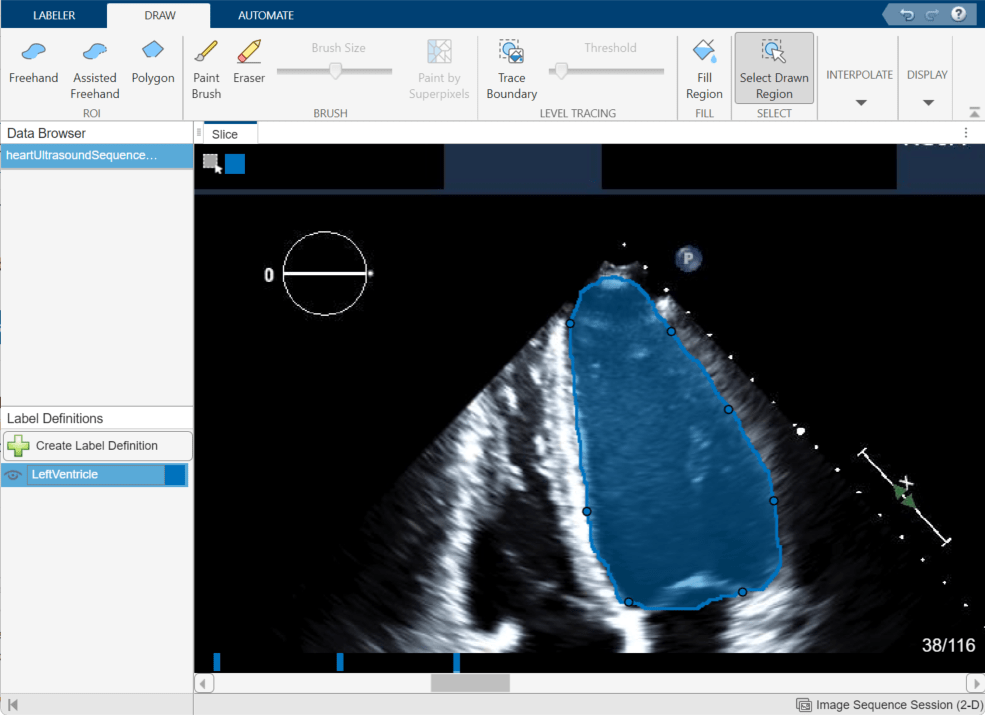
After using interpolation, check the individual frames to see if the interpolation created satisfactory labels. You can manually correct labels using the Paint Brush and Eraser tools. Alternatively, you can refine the labels using one of the tools in the Automate tab. For example, you can use Active Contours to grow an ROI in a frame where it does not fill the full area of the ventricle. You can also undo the interpolation and try interpolating across fewer frames to improve results.
Modify Labels
To refine drawn labels, you can remove label data from individual pixels, from individual frames within an image series, from an entire image, or from the whole labeling session.
Remove labels from individual pixels — In the Draw tab, use the Eraser tool.
Remove labels from one connected region in a slice — Click Select Drawn Region and, in the slice pane, select the region from which you want to remove labels. You can also select the label to remove by right-clicking on the label within the slice plane and selecting Select Drawn Region. Press Delete, or right-click and select Delete, to remove labels from the region.
Remove all labels from an image series — Right-click the file name of the image series in the Data Browser and select Remove Labels. Removing labels from an image removes all pixel labels for all label definitions within the file.
Remove a label from all images within the app session — In the Label Definitions pane, right-click the label name and select Delete. This deletes the label from the
groundTruthMedicalobject in the session folder, and removes all pixel labels for the deleted label name in all images in the session.
Apply Custom Automation Algorithm
You can add a custom automation algorithm to use in the app. On the Automate tab, click Add Algorithm. Import an existing algorithm by selecting From File, or create a new algorithm using the provided function or class template. See Automate Labeling in Medical Image Labeler for an example that applies a custom automation algorithm for 2-D labeling.
Under Slice-Based, select the New option and click Function Template to create a new function that operates on each 2-D image frame. The app opens the template in the MATLAB editor. Replace the sample code in the template with the code for your algorithm. Your function must accept two arguments: each frame as a separate image, and a mask. Your function must also return a mask image.
When you are done editing the template, save the file. The Medical Image Labeler app automatically creates a button in the Automate tab of the toolstrip for your function. To test your function, select it and click Run. By default, the app applies the function to only the current frame.
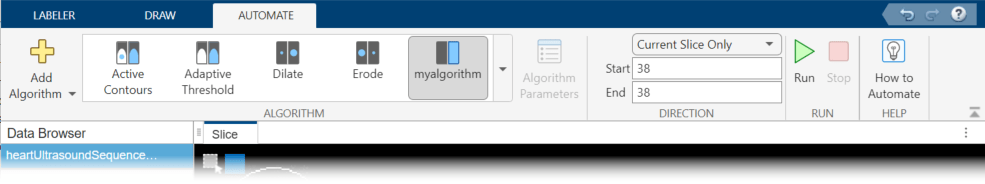
After testing your function on a single frame, you can run it on all of the frames, or a subset of the frames. You can run it from the current frame to the end (the highest numbered frame) or from the current frame back to the beginning (frame 1). You can also specify a range of frames by specifying the starting frame and the ending frame.
Export Ground Truth Data
For this example, the labels for heartUltrasoundSequenceVideo.dcm are complete when you have labeled the ventricle in each frame. The app automatically assigns a value of 0 to unlabeled pixels in the label images saved to the session folder, so you do not need to label the background manually.
The Medical Image Labeler app automatically saves a groundTruthMedical object as a MAT file in the session folder. You can also export a groundTruthMedical object, saved as a MAT file, to an alternate file location from the app. On the Labeler tab, click Export and, under Ground Truth, select To File.
You can load the exported MAT file into the MATLAB workspace using the load function. The properties of the groundTruthMedical object, gTruthMed, contain information about the image data source, label definitions, and location of the saved label images. Display information about the object and each of its properties using these commands.
gTruthMed— Display the properties of thegroundTruthMedicalobject.gTruthMed.DataSource— Location of the source of the unlabeled medical images and image series.gTruthMed.LabelDefinitions— Table of information about label definitions.gTruthMed.LabelData— Locations of the saved, labeled images.
See Also
Medical Image
Labeler | groundTruthMedical