Run Custom Training Loops on a GPU and in Parallel
You can speed up your custom training loops by running them on a GPU, in parallel using multiple GPUs, or on a cluster.
It is recommended to train using a GPU or multiple GPUs. Only use single CPU or multiple CPUs if you do not have a GPU. CPUs are normally much slower than GPUs for both training and inference. Running on a single GPU typically offers much better performance than running on multiple CPU cores.
Note
This topic shows how to perform custom training on GPUs, in parallel, and on the
cloud. To learn about parallel and GPU workflows using the simpler trainnet function, see these topics:
Using a GPU or parallel options requires Parallel Computing Toolbox™. Using a GPU also requires a supported GPU device. For information on supported devices, see GPU Computing Requirements (Parallel Computing Toolbox). Using a remote cluster also requires MATLAB® Parallel Server™.
Train Network on GPU
By default, custom training loops run on the CPU. You can perform automatic
differentiation using dlgradient and dlfeval
on the GPU when your data is on the GPU. To run a custom training loop on a GPU,
convert your data to a gpuArray (Parallel Computing Toolbox) object during training.
You can use minibatchqueue to manage your data during training.
minibatchqueue automatically prepares data for training,
including custom preprocessing and converting data to dlarray
and gpuArray objects. By default, minibatchqueue
returns all mini-batch variables on the GPU if one is available. You can choose
which variables to return on the GPU using the
OutputEnvironment property.
For an example that shows how to use minibatchqueue to train on the GPU, see Train Network Using Custom Training Loop.
Alternatively, you can manually convert your data to a gpuArray
object within the training loop.
To easily specify the execution environment, create the variable executionEnvironment that contains either "cpu", "gpu", or "auto".
executionEnvironment = "auto"During training, after reading a mini-batch, check the execution environment option and
convert the data to a gpuArray if necessary. The canUseGPU
function checks for useable
GPUs.
if (executionEnvironment == "auto" && canUseGPU) || executionEnvironment == "gpu" X = gpuArray(X); end
Train Single Network in Parallel
When you train in parallel, each worker trains the network simultaneously using a portion of a mini-batch. This behavior means that you must combine the gradients, loss, and state parameters after each iteration according to the proportion of the mini-batch processed by each worker.
You can train in parallel on your local machine or on a remote cluster, for example, in the cloud. Start a parallel pool using the desired resources and partition your data between the workers. During training, combine the gradients, loss, and state after each iteration so that the learnable parameters on each worker update in synchronization. For an example that shows how to perform custom training in parallel, see Train Network in Parallel with Custom Training Loop
Set Up Parallel Environment
It is recommended to train using a GPU or multiple GPUs. Only use single CPU or multiple CPUs if you do not have a GPU. CPUs are normally much slower than GPUs for both training and inference. Running on a single GPU typically offers much better performance than running on multiple CPU cores.
Set up the parallel environment before training. Start a parallel pool using the desired resources. To train using multiple GPUs, start a parallel pool with as many workers as available GPUs. MATLAB assigns a different GPU to each worker.
If you are using your local machine, use canUseGPU or gpuDeviceCount (Parallel Computing Toolbox) to determine
whether you have GPUs available. For example, check the availability of your
GPUs and start a parallel pool with as many workers as available GPUs.
if canUseGPU executionEnvironment = "gpu"; numberOfGPUs = gpuDeviceCount("available"); pool = parpool(numberOfGPUs); else executionEnvironment = "cpu"; pool = parpool; end
If you are running code using a remote cluster, for example, a cluster in the cloud, start a parallel pool with as many workers as the number of GPUs per machine multiplied by the number of machines.
For more information on selecting specific GPUs, see Select Particular GPUs to Use for Training.
Specify Mini-Batch Size and Partition Data
Specify the mini-batch size to use during training. For GPU training, scale up the mini-batch size linearly with the number of GPUs to keep the workload on each GPU constant. For example, if you are training on a single GPU using a mini-batch size of 64 and you want to scale up to training with four GPUs of the same type, increase the mini-batch size to 256 so that each GPU processes 64 observations per iteration.
Scale up the mini-batch size by the number of workers, where
N is the number of workers in your parallel pool.
if executionEnvironment == "gpu" miniBatchSize = miniBatchSize.*N end
To use a mini-batch size that is not exactly divisible by the number of workers in your parallel pool, distribute the remainder across the workers.
workerMiniBatchSize = floor(miniBatchSize./repmat(N,1,N)); remainder = miniBatchSize - sum(workerMiniBatchSize); workerMiniBatchSize = workerMiniBatchSize + [ones(1,remainder) zeros(1,N-remainder)]
At the start of training, shuffle the data. Partition the data so that each
worker has access to a portion of the mini-batch. To partition a datastore, use
the partition function.
Use minibatchqueue to manage the data on each worker during training.
A minibatchqueue object automatically prepares data for
training, including custom preprocessing and converting data to
dlarray and gpuArray objects. Create a
minibatchqueue object on each worker using the partitioned
datastore. Set the MiniBatchSize property to the mini-batch
sizes calculated for each worker.
At the start of each training iteration, use the spmdReduce (Parallel Computing Toolbox) function to check that all worker
minibatchqueue objects can return data. If any worker runs
out of data, training stops. If the overall mini-batch size is not exactly
divisible by the number of workers and you do not discard partial mini-batches,
some workers might run out of data before others.
Write your training code inside an spmd (Parallel Computing Toolbox) block so that the
training loop executes on each worker.
% Shuffle the datastore. augimdsTrain = shuffle(augimdsTrain); spmd % Partition the datastore. workerImds = partition(augimdsTrain,N,spmdIndex); % Create a minibatchqueue object using the partitioned datastore on each worker. workerMbq = minibatchqueue(workerImds,... MiniBatchSize = workerMiniBatchSize(spmdIndex),... MiniBatchFcn = @preprocessMiniBatch); ... for epoch = 1:numEpochs % Reset and shuffle the mini-batch queue on each worker. shuffle(workerMbq); % Loop over the mini-batches. while spmdReduce(@and,hasdata(workerMbq)) % Custom training loop ... end ... end end
Aggregate Gradients
To ensure that the network on each worker learns from all the data and not just the data on that worker, aggregate the gradients and use the aggregated gradients to update the network on each worker.
For example, suppose you are training the network net using
the model loss function modelLoss. Your training loop
contains the code for evaluating the loss, gradients, and statistics on each
worker, where workerX and workerT are the
predictor and target response on each worker,
respectively.
[workerLoss,workerGradients,workerState] = dlfeval(@modelLoss,net,workerX,workerT);
To aggregate the gradients, use a weighted sum. Define a helper function to sum the gradients.
function gradients = aggregateGradients(gradients,factor) gradients = extractdata(gradients); gradients = spmdPlus(factor*gradients); end
Inside the training loop, use dlupdate to apply the function to the gradients of each
learnable
parameter.
workerGradients.Value = dlupdate(@aggregateGradients,workerGradients.Value,{workerNormalizationFactor});Aggregate Loss and Accuracy
To find the network loss and accuracy, for example, to plot them during
training to monitor training progress, aggregate the values of the loss and
accuracy on all of the workers. Typically, the aggregated value is the sum of
the value on each worker weighted by the proportion of the mini-batch that each
worker uses. To aggregate the losses and accuracy each iteration, calculate the
weight factor for each worker and use spmdPlus (Parallel Computing Toolbox) to sum the values on each
worker.
workerNormalizationFactor = workerMiniBatchSize(spmdIndex)./miniBatchSize; loss = spmdPlus(workerNormalizationFactor*extractdata(dlworkerLoss)); accuracy = spmdPlus(workerNormalizationFactor*extractdata(dlworkerAccuracy));
Aggregate Statistics
If your network contains layers that track the statistics of your training data, such as batch normalization layers, then you must aggregate the statistics across all workers after each training iteration. Aggregating the statistics ensures that the network learns statistics that are representative of the entire training set.
You can identify the layers that contain statistics before training. For
example, find the relevant layers using a dlnetwork object with
batch normalization
layers.
batchNormLayers = arrayfun(@(l)isa(l,'nnet.cnn.layer.BatchNormalizationLayer'),net.Layers); batchNormLayersNames = string({net.Layers(batchNormLayers).Name}); state = net.State; isBatchNormalizationStateMean = ismember(state.Layer,batchNormLayersNames) & state.Parameter == "TrainedMean"; isBatchNormalizationStateVariance = ismember(state.Layer,batchNormLayersNames) & state.Parameter == "TrainedVariance";
where N is the total number of workers, M is the total number of observations in a mini-batch, mj is the number of observations processed on the jth worker, and are the mean and variance statistics calculated on that worker, respectively, and is the aggregated mean across all workers.
function state = aggregateState(state,factor,... isBatchNormalizationStateMean,isBatchNormalizationStateVariance) stateMeans = state.Value(isBatchNormalizationStateMean); stateVariances = state.Value(isBatchNormalizationStateVariance); for j = 1:numel(stateMeans) meanVal = stateMeans{j}; varVal = stateVariances{j}; % Calculate combined mean. combinedMean = spmdPlus(factor*meanVal); % Calculate combined variance terms to sum. varTerm = factor.*(varVal + (meanVal - combinedMean).^2); % Update state. stateMeans{j} = combinedMean; stateVariances{j} = spmdPlus(varTerm); end state.Value(isBatchNormalizationStateMean) = stateMeans; state.Value(isBatchNormalizationStateVariance) = stateVariances; end
Inside the training loop, use the helper function to update the state of the batch normalization layers with the combined mean and variance.
net.State = aggregateState(workerState,workerNormalizationFactor,...
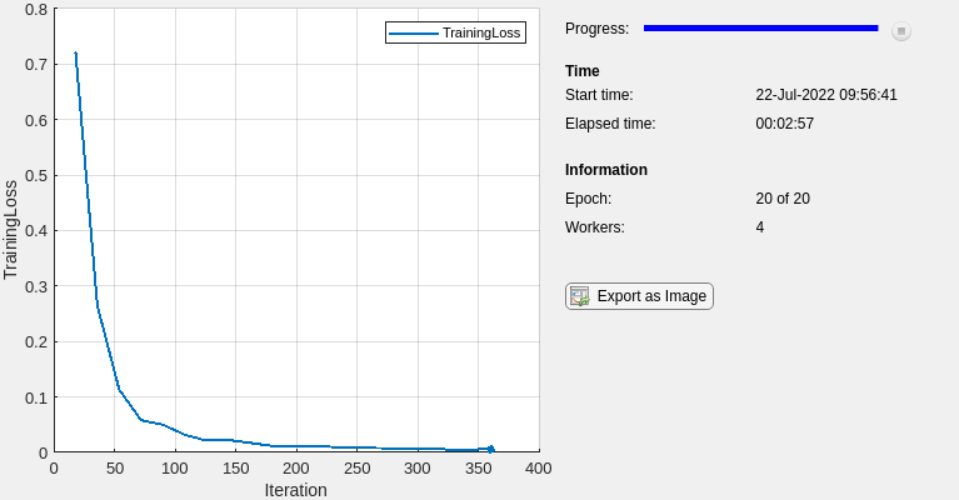
isBatchNormalizationStateMean,isBatchNormalizationStateVariance);Plot Results During Training
To plot results during training, send data from the workers to the client
using a DataQueue (Parallel Computing Toolbox) object.
To plot training progress, set plots to
"training-progress". Otherwise, set
plots to
"none".
plots = "training-progress";Before training perform these steps:
Initialize the
TrainingProgressMonitorobject to track and plot the loss for the network. Because the timer starts when you create the monitor, create the object immediately before the training loop.Initialize a
PollableDataQueueobject for sending a flag to stop training when you click the Stop button. (since R2025a)Before R2025a: Use
spmd(Parallel Computing Toolbox) to initialize aparallel.pool.DataQueue(Parallel Computing Toolbox) object on the workers for sending a flag to stop training when you click the Stop button.Initialize a
DataQueueobject on the client for receiving data from the workers during training.Use
afterEach(Parallel Computing Toolbox) to call thedisplayTrainingProgressfunction each time a worker sends data to the client.
Before R2023a: To plot training progress, create an
animatedline object instead of
initializing a TrainingProgressMonitor object and use the addpoints function inside the
displayTrainingProgress function to update the animatedline.
if plots == "training-progress" % Initialize the training progress monitor. monitor = trainingProgressMonitor( ... Metrics="TrainingLoss", ... Info=["Epoch","Workers"], ... XLabel="Iteration"); % Initialize a PollableDataQueue for sending a stop flag. stopTrainingQueue = parallel.pool.PollableDataQueue(Destination="any"); % Initialize a DataQueue object on the client for sending data from the workers to the client. dataQueue = parallel.pool.DataQueue; % Call displayTrainingProgress each time a worker sends data to the client. displayFcn = @(x) displayTrainingProgress(x,numEpochs,numWorkers,monitor,stopTrainingQueue); afterEach(dataQueue,displayFcn) end
displayTrainingProgress helper function
updates the Training Progress window and checks whether the
Stop button has been clicked. If you click the
Stop button the PollableDataQueue
object instructs the workers to stop
training.function displayTrainingProgress(data,numEpochs,numWorkers,monitor,stopTrainingQueue) % Extract epoch, iteration, and loss data. epoch = data(1); iteration = data(2); loss = data(3); % Update the training progress monitor. recordMetrics(monitor,iteration,TrainingLoss=loss); updateInfo(monitor,Epoch=epoch + " of " + numEpochs,Workers=numWorkers); monitor.Progress = 100*epoch/numEpochs; % Send a flag to the workers if the Stop button has been clicked. if monitor.Stop send(stopTrainingQueue,true); end end
Inside the training loop, at the end of each iteration or epoch, check whether
the Stop button has been clicked and use the
DataQueue object to send the training data from the workers
to the client. At the end of each iteration, the aggregated loss is the same on
each worker, so you can send data from a single
worker.
spmd epoch = 0; iteration = 0; stopRequest = false; % Prepare input data and mini-batches. ... % Loop over epochs. while epoch < numEpochs && ~stopRequest epoch = epoch + 1; % Reset and shuffle the mini-batch queue on each worker. ... % Loop over mini-batches. while spmdReduce(@and,hasdata(workerMbq)) && ~stopRequest iteration = iteration + 1; % Custom training loop. ... if plots == "training-progress" % Check whether the Stop button has been clicked. stopRequest = spmdPlus(stopTrainingEventQueue.QueueLength); % Send training progress information to the client. if spmdIndex == 1 data = [epoch iteration loss]; send(dataQueue,gather(data)); end end end end end
Train Multiple Networks in Parallel
To train multiple networks in parallel, start a parallel pool and use parfor (Parallel Computing Toolbox) to train a single network
on each worker.
You can run the training locally or on a remote cluster. Using a remote cluster requires a MATLAB Parallel Server license. For more information about managing cluster resources, see Discover Clusters and Use Cluster Profiles (Parallel Computing Toolbox). If you have multiple GPUs and want to exclude some from training, you can choose to train on only some GPUs. For more information on selecting specific GPUs, see Select Particular GPUs to Use for Training.
You can modify the network or training parameters on each worker to perform
parameter sweeps in parallel. For example, if networks is an
array of dlnetwork objects, you can use this code to train multiple
different networks using the same data. After the parfor-loop
finishes, trainedNetworks contains the resulting networks trained
by the
workers.
parpool; parfor idx = 1:numNetworks iteration = 0; velocity = []; % Allocate one network per worker. net = networks(idx) % Loop over epochs. for epoch = 1:numEpochs % Shuffle data. shuffle(mbq); % Loop over mini-batches. while hasdata(mbq) iteration = iteration + 1; % Custom training loop. ... end end % Send the trained networks back to the client. trainedNetworks{idx} = net; end
Use Experiment Manager to Train in Parallel
You can use Experiment Manager to run your custom training loops in parallel. You can run multiple trials simultaneously or run a single trial at a time using parallel resources.
To run multiple trials at the same time using one parallel worker for each trial,
set up your custom training experiment and set Mode to
Simultaneous before running your experiment.
To run a single trial at a time using multiple parallel workers, define your
parallel environment in your experiment training function, use an
spmd block to train the network in parallel, and set
Mode to Sequential. For more information
on training a single network in parallel with a custom training loop, see Train Single Network in Parallel and Custom Training with Multiple GPUs in Experiment Manager.
To display the training plot and track the progress of each trial while the experiment is running, under Review Results, click Training Plot.
For more information about training in parallel using Experiment Manager, see Run Experiments in Parallel.
See Also
spmd (Parallel Computing Toolbox) | parfor (Parallel Computing Toolbox) | TrainingProgressMonitor | gpuArray (Parallel Computing Toolbox) | dlarray | dlnetwork | deep.gpu.deterministicAlgorithms