bioinfo.pipeline.block.FileChooser
Description
A FileChooser block enables you to select files or download files
from URLs.
Creation
Syntax
Description
Input Arguments
Names of files or URLs, specified as a string, character vector, string vector, or cell array of character vectors. You can include a full file path. The block does not use the MATLAB® path.
Data Types: char | string | cell
Properties
Function to handle errors from the run
method of the block, specified as a function handle. The handle specifies the function to call
if the run method encounters an error within a pipeline. For the pipeline to continue after a
block fails, ErrorHandler must return a structure that is compatible with
the output ports of the block. The error handling function is called with the following two inputs:
Structure with these fields:
Field Description identifier Identifier of the error that occurred message Text of the error message index Linear index indicating which block process failed in the parallel run. By default, the index is 1 because there is only one run per block. For details on how block inputs can be split across different dimensions for multiple run calls, see Bioinformatics Pipeline SplitDimension. Input structure passed to the
runmethod when it fails
Data Types: function_handle
Names of files or URLs, specified as a string, character vector, string vector, or cell array of character vectors.
Files is always appended after PathRoot to
determine the file or URL destinations. Files can include file,
http, https as a scheme if
PathRoot is empty.
Data Types: char | string | cell
This property is read-only.
Input ports of the block, specified as a structure. The field
names of the structure are the names of the block input ports, and the field values are bioinfo.pipeline.Input objects. These objects describe the input port behaviors.
The input port names are the expected field names of the input structure that you pass to the
block run method.
Data Types: struct
This property is read-only.
Output ports of the block, specified as a structure. The field
names of the structure are the names of the block output ports, and the field values are bioinfo.pipeline.Output objects. These objects describe the output port behaviors.
The field names of the output structure returned by the block run method
are the same as the output port names.
The FileChooser block Outputs structure has
the following field:
Data Types: struct
Parameters for obtaining data from a web server, specified as a weboptions object.
This property is used only when the scheme is http or
https, or when Files contains a URL. The
default value is a weboptions object with default property
values.
Root path for all files in the Files property, specified as a
string or character vector.
PathRoot is always prefixed to Files to
determine the file or URL destinations.
Data Types: char | string
Object Functions
compile | Perform block-specific additional checks and validations |
copy | Copy array of handle objects |
emptyInputs | Create input structure for use with run method |
eval | Evaluate block object |
run | Run block object |
Examples
Import the Pipeline and block objects needed for the example.
import bioinfo.pipeline.Pipeline import bioinfo.pipeline.block.*
Create a pipeline.
qcpipeline = Pipeline;
Select an input FASTQ file using a FileChooser block.
fastqfile = FileChooser(which("SRR005164_1_50.fastq"));Create a SeqFilter block.
sequencefilter = SeqFilter;
Define the filtering threshold value. Specifically, filter out sequences with a total of more than 10 low-quality bases, where a base is considered a low-quality base if its quality score is less than 20.
sequencefilter.Options.Threshold = [10 20];
Add the blocks to the pipeline.
addBlock(qcpipeline,[fastqfile,sequencefilter]);
Connect the output of the first block to the input of the second block. To do so, you need to first check the input and output port names of the corresponding blocks.
View the Outputs (port of the first block) and Inputs (port of the second block).
fastqfile.Outputs
ans = struct with fields:
Files: [1×1 bioinfo.pipeline.Output]
sequencefilter.Inputs
ans = struct with fields:
FASTQFiles: [1×1 bioinfo.pipeline.Input]
Connect the Files output port of the fastqfile block to the FASTQFiles port of sequencefilter block.
connect(qcpipeline,fastqfile,sequencefilter,["Files","FASTQFiles"]);
Next, create a UserFunction block that calls the seqqcplot function to plot the quality data of the filtered sequence data. In this case, inputFile is the required argument for the seqqcplot function. The required argument name can be anything as long as it is a valid variable name.
qcplot = UserFunction("seqqcplot",RequiredArguments="inputFile",OutputArguments="figureHandle");
Alternatively, you can also use dot notation to set up your UserFunction block.
qcplot = UserFunction; qcplot.RequiredArguments = "inputFile"; qcplot.Function = "seqqcplot"; qcplot.OutputArguments = "figureHandle";
Add the block.
addBlock(qcpipeline,qcplot);
Check the port names of sequencefilter block and qcplot block.
sequencefilter.Outputs
ans = struct with fields:
FilteredFASTQFiles: [1×1 bioinfo.pipeline.Output]
NumFilteredIn: [1×1 bioinfo.pipeline.Output]
NumFilteredOut: [1×1 bioinfo.pipeline.Output]
qcplot.Inputs
ans = struct with fields:
inputFile: [1×1 bioinfo.pipeline.Input]
Connect the FilteredFASTQFiles port of the sequencefilter block to the inputFile port of the qcplot block.
connect(qcpipeline,sequencefilter,qcplot,["FilteredFASTQFiles","inputFile"]);
Run the pipeline to plot the sequence quality data.
run(qcpipeline);
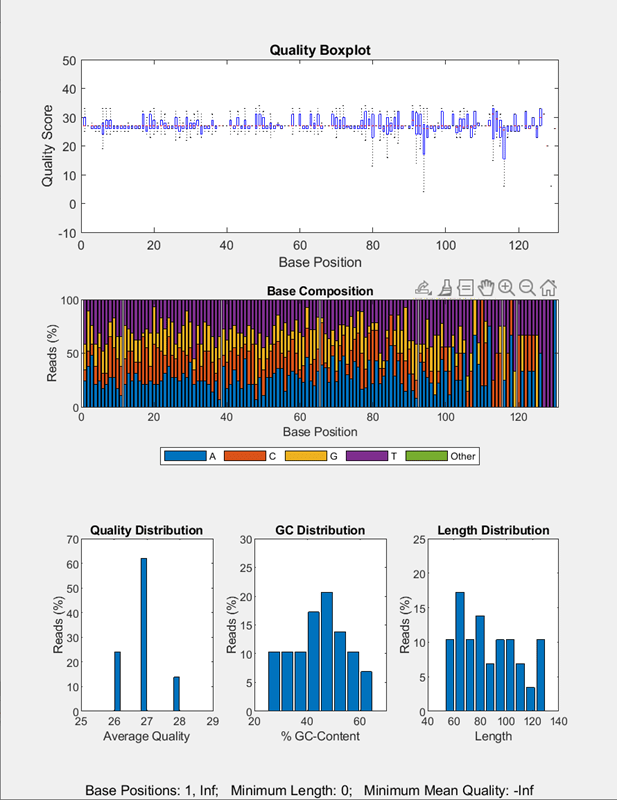
Use a FileChooser block to select an input file
provided with the toolbox.
import bioinfo.pipeline.block.FileChooser import bioinfo.pipeline.Pipeline FC = FileChooser(which("SRR6008575_10k_1.fq")); P = Pipeline; addBlock(P, FC); run(P); R = results(P, FC)
R =
struct with fields:
Files: [1×1 bioinfo.pipeline.datatypes.File]Call unwrap on Files to see the location of
the file.
unwrap(R.Files)
ans =
"C:\Program Files\MATLAB\R2023a\toolbox\bioinfo\bioinfodata\SRR6008575_10k_1.fq"Version History
Introduced in R2023a
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
웹사이트 선택
번역된 콘텐츠를 보고 지역별 이벤트와 혜택을 살펴보려면 웹사이트를 선택하십시오. 현재 계신 지역에 따라 다음 웹사이트를 권장합니다:
또한 다음 목록에서 웹사이트를 선택하실 수도 있습니다.
사이트 성능 최적화 방법
최고의 사이트 성능을 위해 중국 사이트(중국어 또는 영어)를 선택하십시오. 현재 계신 지역에서는 다른 국가의 MathWorks 사이트 방문이 최적화되지 않았습니다.
미주
- América Latina (Español)
- Canada (English)
- United States (English)
유럽
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)