smoteTabularSynthesizer
Description
To generate synthetic data, you can first create a
smoteTabularSynthesizer object using an existing multivariate data set.
Then, use the synthesizeTabularData object function to synthesize data using SMOTE (synthetic
minority oversampling technique). After you synthesize data, you can test whether the new data
set comes from the same distribution as the original data set. Use the mmdtest or
knntest function to
determine how close the data distributions are to each other. For more information about
SMOTE, see Generate Synthetic Data Using SMOTE.
Creation
Syntax
Description
synthesizer = smoteTabularSynthesizer(X)synthesizer) using the
existing data X.
synthesizer = smoteTabularSynthesizer(___,Name=Value)
Input Arguments
Existing data set, specified as a numeric matrix or a table. Rows of
X correspond to observations, and columns of
X correspond to variables. Multicolumn variables and cell
arrays other than cell arrays of character vectors are not supported.
Data Types: single | double | table
Name of the class labels variable in X, specified as a
character vector or string scalar. X must be a table, and
Yname must specify a column in X that is a
numeric, categorical, or logical vector; a character or string array; or a cell array
of character vectors. The Yname variable must contain class
labels for one or two classes only.
Data Types: char | string
Class labels for one or two classes, specified as a numeric, categorical, or
logical vector; a character or string array; or a cell array of character vectors.
Rows of Y correspond to observations.
Data Types: single | double | logical | char | string | cell | categorical
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: smoteTabularSynthesizer(X,Y,NumNeighbors=3,Distance="fastseuclidean")
specifies to use 3 nearest neighbors and the fast standardized Euclidean
distance.
Number of nearest neighbors to use when generating synthetic data, specified as a positive integer scalar.
If you do not specify
YnameorY, the default value forNumNeighborsismin(5,size(X,1)).If you specify
YnameorY, the default value forNumNeighborsis a scalar or two-element vector. Each entry ismin(5,n), wherenis the number of observations in the corresponding class.
Example: NumNeighbors=10
Data Types: single | double
Distance metric for finding nearest neighbors, specified as a character vector or string scalar.
If all the variables are continuous (numeric), then you can specify one of the following distance metrics.
Value Description "euclidean"Euclidean distance
"fasteuclidean"Euclidean distance computed by using an alternative algorithm that saves time when the number of variables is at least 10. In some cases, this faster algorithm can reduce accuracy. "seuclidean"Standardized Euclidean distance. Each coordinate difference between observations is scaled by dividing by the corresponding variable standard deviation.
"fastseuclidean"Standardized Euclidean distance computed by using an alternative algorithm that saves time when the number of variables is at least 10. In some cases, this faster algorithm can reduce accuracy. Note
If you specify one of these distance metrics and the data includes categorical variables, then the software treats each categorical variable as a numeric variable for the distance computation, with each category represented by a positive integer.
If all the variables are categorical, then you can specify the following distance metric.
Value Description "hamming"Hamming distance, which is the percentage of coordinates that differ
Note
If you specify this distance metric and the data includes continuous (numeric) variables, then the software treats each continuous variable as a categorical variable for the distance computation.
If the variables are a mix of continuous (numeric) and categorical variables, then you can specify the following distance metric.
Value Description "goodall3"Modified Goodall distance
The default value is "seuclidean" if all the variables are
continuous, "hamming" if all the variables are categorical, and
"goodall3" if the variables are a mix of continuous and
categorical variables.
Example: Distance="euclidean"
Data Types: char | string
Size in megabytes of the cache allocated for the distance computation, specified as
"maximal" or a positive scalar. If the cache size is
"maximal", the software tries to allocate enough memory for an
intermediate matrix.
The CacheSize name-value argument is valid only when the
Distance value is "fasteuclidean",
"fastseuclidean", or "goodall3".
For the
fastdistance metrics, the intermediate matrix corresponds to the Gram matrix.For the modified Goodall distance metric, the intermediate matrix corresponds to the distance matrix.
Example: CacheSize="maximal"
Data Types: single | double | char | string
Variable names, specified as a string array or a cell array of character vectors.
If
Xis a numeric matrix, then you can useVariableNamesto assign names to the variables inX.The order of the names in
VariableNamesmust correspond to the order of the variables inX. That is,VariableNames{1}is the name ofX(:,1),VariableNames{2}is the name ofX(:,2), and so on.size(X,2)andnumel(VariableNames)must be equal.By default,
VariableNamesis{'x1','x2',...}.
If
Xis a table, then you can useVariableNamesto specify which variables to use. That is,smoteTabularSynthesizeruses only the variables inVariableNamesto generate synthetic data.VariableNamesmust be a subset ofX.Properties.VariableNames.By default,
VariableNamescontains the names of all variables, excluding the class labels variableYname.
Example: VariableNames=["SepalLength","SepalWidth","PetalLength","PetalWidth"]
Data Types: string | cell
List of the categorical variables, excluding the class labels variable
Yname, specified as one of the values in this table.
| Value | Description |
|---|---|
| Positive integer vector | Each entry in the vector is an index value indicating that the
corresponding variable is categorical. The index values are between 1 and
v, where v is the number of variables
listed in |
| Logical vector | A |
| String array or cell array of character vectors | Each element in the array is the name of a categorical variable. The names must
match the entries in VariableNames. |
"all" | All variables are categorical. |
By default, if the variables are in a numeric matrix, the software assumes all the variables
are continuous. If the variables are in a table, the software assumes they are
categorical if they are logical vectors, categorical vectors, character
arrays, string arrays, or cell arrays of character vectors. To identify any other
variables as categorical, specify them by using the
CategoricalVariables name-value argument.
Do not specify discrete numeric variables as categorical variables. Use the
DiscreteNumericVariables name-value argument instead.
Example: CategoricalVariables="all"
Data Types: single | double | logical | string | cell
List of the discrete numeric variables, specified as one of the values in this table.
| Value | Description |
|---|---|
| Positive integer vector | Each entry in the vector is an index value indicating that
the corresponding variable is a discrete numeric variable. The
index values are between 1 and v, where
v is the number of variables listed in
|
| Logical vector | A |
| String array or cell array of character vectors | Each element in the array is the name of a discrete numeric
variable. The names must match the entries in
VariableNames. |
"all" | All variables are discrete numeric variables. |
You cannot specify categorical variables as discrete numeric variables.
Example: DiscreteNumericVariables=[2 5]
Data Types: single | double | logical | string | cell
Properties
This property is read-only.
Variable names, returned as a string array. The order of the elements of
VariableNames corresponds to the order in which the variable
names appear in the existing data set X.
Data Types: string
This property is read-only.
Indices of the categorical variables, returned as a positive integer vector. Each
index value in CategoricalVariables indicates that the
corresponding variable listed in VariableNames
is categorical. If none of the variables are categorical, then this property is empty
([]).
Data Types: double
This property is read-only.
Indices of the discrete numeric variables, returned as a positive integer vector.
Each index value in DiscreteNumericVariables indicates that the
corresponding variable listed in VariableNames
is a discrete numeric variable. If none of the variables are discrete numeric variables,
then this property is empty ([]).
Data Types: double
This property is read-only.
Number of nearest neighbors to use when generating synthetic data, returned as a
positive integer scalar or a two-element positive integer vector. If you specify class
labels when you create the synthesizer object, then NumNeighbors(i)
corresponds to the number of nearest neighbors to use for class
ClassNames(i,:).
Data Types: single | double
This property is read-only.
Distance metric used for finding nearest neighbors, returned as
"euclidean", "fasteuclidean",
"fastseuclidean", "goodall3",
"hamming", or "seuclidean". For more information
on these distance metrics, see the Distance
name-value argument.
Data Types: string
This property is read-only.
Names of the classes in Yname or
Y for which
to generate synthetic data, returned as a numeric, categorical, or logical vector; a
character or string array; or a cell array of character vectors. If you do not specify
class labels when you create the synthesizer object, then this property is empty
([]).
Data Types: single | double | logical | char | string | cell | categorical
This property is read-only.
Number of observations in the existing data set X, returned
as a positive integer scalar or a two-element positive integer vector. If you specify
class labels when you create the synthesizer object, then
NumObservations(i) corresponds to the number of observations in the
existing data set that belong to class ClassNames(i,:).
Data Types: double
Object Functions
synthesizeTabularData | Synthesize tabular data using binning-based or SMOTE-based synthesizer |
Examples
Use existing training data to create a smoteTabularSynthesizer object. Then, synthesize data using the synthesizeTabularData object function. Train a model using the existing training data, and then train the same type of model using both the existing training data and the synthetic data. Compare the performance of the two models using test data.
Load the carbig data set, which contains measurements of cars made in the 1970s and early 1980s. Categorize the cars based on whether they were made in Europe.
load carbig Origin = categorical(cellstr(Origin)); Origin = mergecats(Origin,["France","Germany", ... "Sweden","Italy","England"],"Europe"); Origin = mergecats(Origin,["USA","Japan"],"NotEurope"); tabulate(Origin)
Value Count Percent
Europe 73 17.98%
NotEurope 333 82.02%
The data is imbalanced, with only about 18% of cars originating in Europe.
Create a table containing the variables Acceleration, Displacement, and so on, as well as the response variable Origin. Remove rows of cars where the table has missing values.
cars = table(Acceleration,Displacement,Horsepower, ...
MPG,Weight,Origin);
cars = rmmissing(cars);Partition the data into training and test sets. Use approximately 50% of the observations for model training and synthesizing new data, and 50% of the observations for model testing. Use stratified partitioning so that approximately the same ratio of European to non-European cars exists in both the training and test sets.
rng("default")
cv = cvpartition(cars.Origin,Holdout=0.5);
trainCars = cars(training(cv),:);
testCars = cars(test(cv),:);Create a smoteTabularSynthesizer object using the trainCars data set. Specify Origin as the class labels variable.
synthesizer = smoteTabularSynthesizer(trainCars,"Origin")synthesizer =
smoteTabularSynthesizer
VariableNames: ["Acceleration" "Displacement" "Horsepower" "MPG" "Weight"]
ClassNames: [Europe NotEurope]
NumNeighbors: [5 5]
Distance: "seuclidean"
NumObservations: [34 162]
Properties, Methods
synthesizer is a smoteTabularSynthesizer object with two classes (Europe and NotEurope).
Synthesize new data by using synthesizer. Specify to generate 40 observations belonging to the class of European cars only.
syntheticCars = synthesizeTabularData(synthesizer,40,ClassNames="Europe"); The synthesizeTabularData object function uses the information stored in synthesizer to generate syntheticCars.
To visualize the difference between the existing European car data and the synthetic European car data, you can use the detectdrift function. Filter the trainCars data to include European car data only. The detectdrift function uses permutation testing to detect drift between europeanCars and syntheticCars.
europeanCars = trainCars(trainCars.Origin=="Europe",:);
dd = detectdrift(europeanCars,syntheticCars);dd is a DriftDiagnostics object with a plotEmpiricalCDF object function for visualization.
For the continuous variables, use the plotEmpiricalCDF function to see the difference between the empirical cumulative distribution function (ecdf) of the values in europeanCars and the ecdf of the values in syntheticCars.
continuousVariable ="Acceleration"; plotEmpiricalCDF(dd,Variable=continuousVariable) legend(["Real data","Synthetic data"])
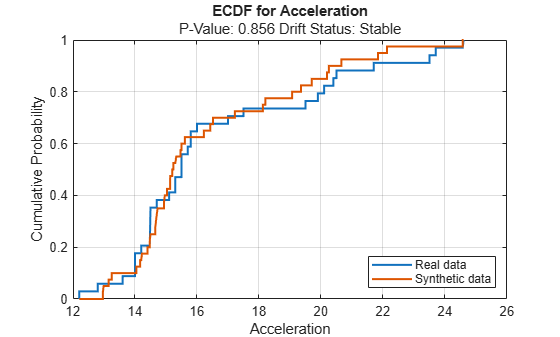
For the Acceleration predictor, the ecdf plot for the existing values (in blue) matches the ecdf plot for the synthetic values (in red) fairly well.
Train an SVM classifier using the original training data trainCars. Specify Origin as the response variable, and standardize the predictors before training. Then, train the same kind of classifier using both the original data and the synthetic data (syntheticCars).
originalMdl = fitcsvm(trainCars,"Origin",Standardize=true); newMdl = fitcsvm([trainCars;syntheticCars],"Origin",Standardize=true);
Evaluate the performance of the two models on the test set using confusion matrices.
originalPredictions = predict(originalMdl,testCars); newPredictions = predict(newMdl,testCars); tiledlayout(1,2) nexttile confusionchart(testCars.Origin,originalPredictions) title("Original Model") nexttile confusionchart(testCars.Origin,newPredictions) title("New Model")
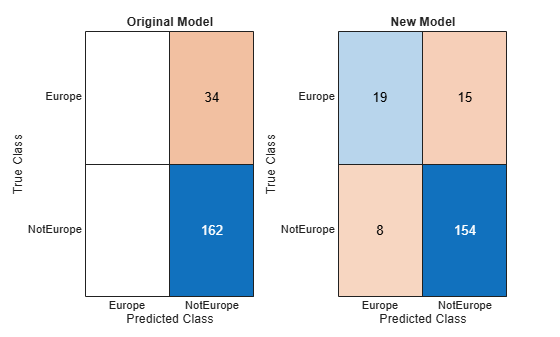
The model trained on the original data classifies all test observations as non-European cars. The model trained on the original and synthetic data has greater accuracy than the other model and correctly classifies the majority of European cars in the test set.
Evaluate data synthesized from an existing data set. Compare the existing and synthetic data sets for each class to determine the similarity between the two multivariate data distributions.
Load the sample file fisheriris.csv, which contains iris data including sepal length, sepal width, petal width, and species type. Read the file into a table, and then display the first eight observations in the table.
fisheriris = readtable("fisheriris.csv");
head(fisheriris) SepalLength SepalWidth PetalLength PetalWidth Species
___________ __________ ___________ __________ __________
5.1 3.5 1.4 0.2 {'setosa'}
4.9 3 1.4 0.2 {'setosa'}
4.7 3.2 1.3 0.2 {'setosa'}
4.6 3.1 1.5 0.2 {'setosa'}
5 3.6 1.4 0.2 {'setosa'}
5.4 3.9 1.7 0.4 {'setosa'}
4.6 3.4 1.4 0.3 {'setosa'}
5 3.4 1.5 0.2 {'setosa'}
Display the types of irises in the table and their proportion in the data set.
tabulate(fisheriris.Species)
Value Count Percent
setosa 50 33.33%
versicolor 50 33.33%
virginica 50 33.33%
The data set contains three types of irises (setosa, versicolor, and virginica) with 50 observations each.
Create separate tables for the versicolor data and the virginica data. Combine the two tables into one.
versicolorData = fisheriris(fisheriris.Species=="versicolor",:); virginicaData = fisheriris(fisheriris.Species=="virginica",:); irisData = [versicolorData;virginicaData]
irisData=100×5 table
SepalLength SepalWidth PetalLength PetalWidth Species
___________ __________ ___________ __________ ______________
7 3.2 4.7 1.4 {'versicolor'}
6.4 3.2 4.5 1.5 {'versicolor'}
6.9 3.1 4.9 1.5 {'versicolor'}
5.5 2.3 4 1.3 {'versicolor'}
6.5 2.8 4.6 1.5 {'versicolor'}
5.7 2.8 4.5 1.3 {'versicolor'}
6.3 3.3 4.7 1.6 {'versicolor'}
4.9 2.4 3.3 1 {'versicolor'}
6.6 2.9 4.6 1.3 {'versicolor'}
5.2 2.7 3.9 1.4 {'versicolor'}
5 2 3.5 1 {'versicolor'}
5.9 3 4.2 1.5 {'versicolor'}
6 2.2 4 1 {'versicolor'}
6.1 2.9 4.7 1.4 {'versicolor'}
5.6 2.9 3.6 1.3 {'versicolor'}
6.7 3.1 4.4 1.4 {'versicolor'}
⋮
Create 200 new observations from the data in irisData: 100 for the versicolor class and 100 for the virginica class. First, create an object by using the smoteTabularSynthesizer function. Then, synthesize the data by using the synthesizeTabularData object function.
To better compare class-specific data later on, call the synthesizeTabularData function twice and return versicolor and virginica synthetic data separately.
synthesizer = smoteTabularSynthesizer(irisData,"Species"); syntheticVersicolorData = synthesizeTabularData(synthesizer,100, ... ClassNames="versicolor"); syntheticVirginicaData = synthesizeTabularData(synthesizer,100, ... ClassNames="virginica");
For each type of iris, visually compare the observations in the existing data set and the synthetic data set by using scatter plots. Each point corresponds to an observation. The point color indicates the species of the corresponding iris (blue for versicolor and red for virginica).
tiledlayout(2,2) nexttile scatter(versicolorData,"SepalLength","PetalLength") title("Existing Versicolor Data") nexttile scatter(syntheticVersicolorData,"SepalLength","PetalLength") title("Synthetic Versicolor Data") nexttile scatter(virginicaData,"SepalLength","PetalLength", ... MarkerEdgeColor="red") title("Existing Virginica Data") nexttile scatter(syntheticVirginicaData,"SepalLength","PetalLength", ... MarkerEdgeColor="red") title("Synthetic Virginica Data")
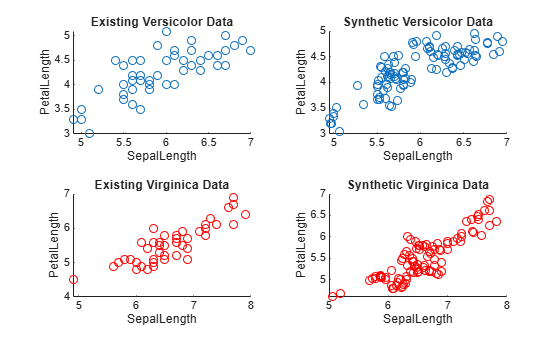
For each iris type, the scatter plots indicate that the existing data set and the synthetic data set have similar characteristics.
For each iris type, compare the existing and synthetic data sets by using the knntest function. The function performs a two-sample hypothesis test for the null hypothesis that the data sets come from the same distribution.
[knnstat1,p1,h1] = knntest(versicolorData,syntheticVersicolorData)
knnstat1 = 0.5327
p1 = 0.8830
h1 = 0
[knnstat2,p2,h2] = knntest(virginicaData,syntheticVirginicaData)
knnstat2 = 0.5400
p2 = 0.7738
h2 = 0
For each iris type, the returned value of h = 0 indicates that knntest fails to reject the null hypothesis that the existing data set and the synthetic data set come from different distributions at the 5% significance level. As with other hypothesis tests, this result does not guarantee that the null hypothesis is true. That is, the data sets do not necessarily come from the same distribution, but the high p-value indicates that the distributions of the existing and synthetic data sets are similar.
Algorithms
SMOTE (synthetic minority oversampling technique) is a technique for generating synthetic data when you have an imbalanced data set, that is, when the number of observations is not uniform across the response classes.
Assume the data set X has two classes, where class kbig has many more observations than class ksmall. If the data set X has only one class, then assume
all observations belong to ksmall. When you use SMOTE, the synthesizeTabularData
function generates each new ksmall observation in the following way:
Randomly select an observation x in ksmall.
In the data set, find the
NumNeighbors-nearest neighbors of x that also belong to ksmall.Randomly select one of the nearest neighbors .
For each continuous predictor p, set the predictor value of the new observation to , where xp is the predictor value of the original observation x, is the predictor value of the nearest neighbor , and r is a random value in (0,1) as selected by the
randfunction.For the categorical predictors q1, …, qm, set the predictor values of the new observation to the mode of the vector of predictor values among the
NumNeighbors-nearest neighbors of x. In the case of a tie, choose a vector at random.
The function follows the same process to generate observations in class kbig. The number of synthetic observations generated depends on
n (the input argument of
synthesizeTabularData).
References
[1] Chawla, Nitesh V., Kevin W. Bowyer, Lawrence O. Hall, and W. Philip Kegelmeyer. "SMOTE: Synthetic Minority Over-sampling Technique." Journal of Artificial Intelligence Research 16 (2002): 321-357.
Version History
Introduced in R2026a
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
웹사이트 선택
번역된 콘텐츠를 보고 지역별 이벤트와 혜택을 살펴보려면 웹사이트를 선택하십시오. 현재 계신 지역에 따라 다음 웹사이트를 권장합니다:
또한 다음 목록에서 웹사이트를 선택하실 수도 있습니다.
사이트 성능 최적화 방법
최고의 사이트 성능을 위해 중국 사이트(중국어 또는 영어)를 선택하십시오. 현재 계신 지역에서는 다른 국가의 MathWorks 사이트 방문이 최적화되지 않았습니다.
미주
- América Latina (Español)
- Canada (English)
- United States (English)
유럽
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)