plotResiduals
Plot residuals of linear mixed-effects model
Syntax
Description
plotResiduals(
specifies additional options using one or more name-value arguments. For
example, you can specify the residual type and the graphical properties of
residual data points.lme,plottype,Name=Value)
h = plotResiduals(___)h to the lines or patches in the plot of
residuals.
Examples
Load the sample data.
load("weight.mat")weight contains data from a longitudinal study, where 20 subjects are randomly assigned to 4 exercise programs, and their weight loss is recorded over six 2-week time periods. This is simulated data.
Store the data in a table. Define Subject and Program as categorical variables.
tbl = table(InitialWeight,Program,Subject,Week,y); tbl.Subject = categorical(tbl.Subject); tbl.Program = categorical(tbl.Program);
Fit a linear mixed-effects model where the initial weight, type of program, week, and the interaction between the week and type of program are the fixed effects. The intercept and week vary by subject.
lme = fitlme(tbl,"y ~ InitialWeight + Program*Week + (Week|Subject)");Plot the histogram of the raw residuals.
plotResiduals(lme)
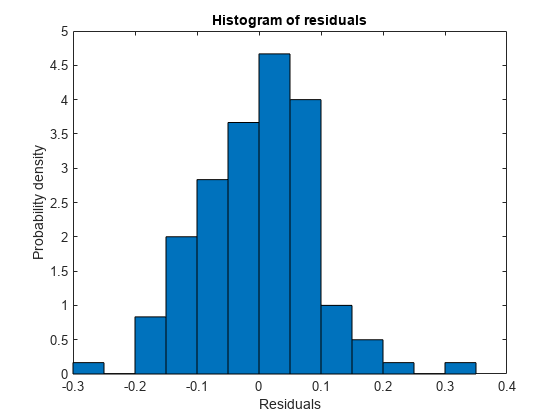
Plot the residuals versus the fitted values.
plotResiduals(lme,"fitted")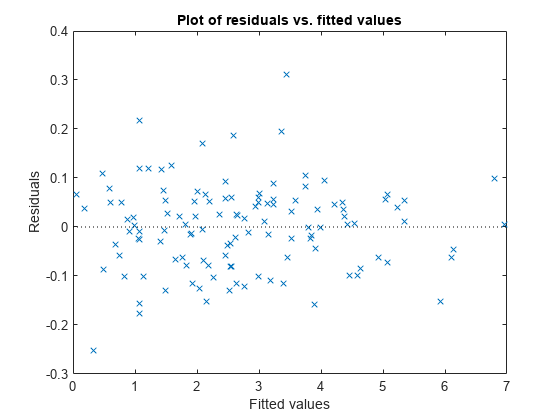
There is no obvious pattern, so there are no immediate signs of heteroscedasticity.
Create the normal probability plot of residuals.
plotResiduals(lme,"probability")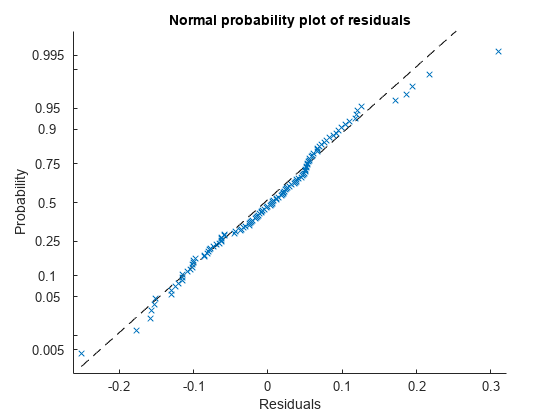
Data appears to be normal.
Find the observation number for the data that appears to be an outlier to the right of the plot.
find(residuals(lme)>0.25)
ans = 101
Create a box plot of the raw, Pearson, and standardized residuals.
r = residuals(lme); pr = residuals(lme,ResidualType="Pearson"); st = residuals(lme,ResidualType="Standardized"); X = [r pr st]; boxplot(X,labels=["Raw","Pearson","Standardized"])
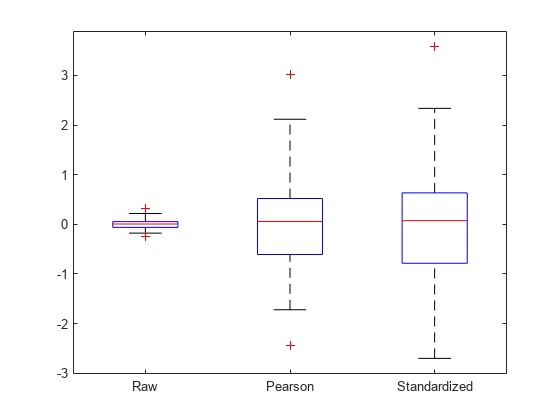
All three box plots point out the outlier on the right tail of the distribution. The box plots of raw and Pearson residuals also point out a second possible outlier on the left tail. Find the corresponding observation number.
find(pr<-2)
ans = 10
Plot the raw residuals versus lagged residuals.
plotResiduals(lme,"lagged")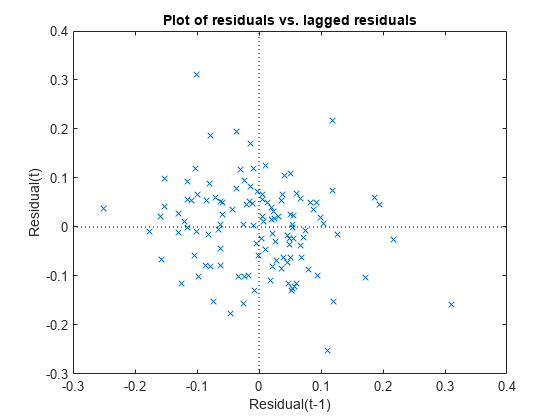
There is no obvious pattern in the graph. The residuals do not appear to be correlated.
Input Arguments
Linear mixed-effects model, specified as a LinearMixedModel object constructed using fitlme or fitlmematrix.
Type of residual plot, specified as one of the following.
"histogram" | Default. Histogram of residuals |
"caseorder" | Residuals versus case (row) order |
"fitted" | Residuals versus fitted values |
"lagged" | Residuals versus lagged residual (r(t) versus r(t – 1)) |
"probability" | Normal probability plot |
"observed" | Observed vs. fitted values. This plot includes a dotted reference line of y = x. Each residual is represented by the vertical distance from the corresponding observed value to the reference line. |
"symmetry" | Symmetry plot |
Example: plotResiduals(lme,"lagged")
Since R2024a
Target axes, specified as an Axes object. If you do not specify the axes,
then plotResiduals uses the current axes (gca).
Name-Value Arguments
Specify optional pairs of arguments as
Name1=Value1,...,NameN=ValueN, where Name is
the argument name and Value is the corresponding value.
Name-value arguments must appear after other arguments, but the order of the
pairs does not matter.
Example: plotResiduals(lme,"probability",ResidualType="Standardized",Marker="o")
specifies the residual type as standardized and the marker symbol as
o for a probability plot.
The plotResiduals name-value arguments listed here are only a
subset. For a complete list, see Patch Properties when the plot type is
"histogram" and Line Properties for all other plot types.
Residual type, specified as one of the following.
| Residual Type | Conditional | Marginal |
|---|---|---|
"Raw" |
|
|
"Pearson" |
|
|
"Standardized" |
|
|
Example: ResidualType="Standardized"
Data Types: string | char
Histogram face color, specified as "interp",
"flat" an RGB triplet, a hexadecimal color code,
a color name, or a short name. FaceColor is
supported only when plottype is
"histogram".
To create a different color for each face, specify the
CData or FaceVertexCData
property as an array containing one color per face or one color per
vertex. The colors can be interpolated from the colors of the
surrounding vertices of each face, or they can be uniform. For
interpolated colors, specify this property as
"interp". For uniform colors, specify this
property as "flat". If you specify
"flat" and a different color for each vertex, the
color of the first vertex you specify determines the face color.
To designate a single color for all of the faces, specify this property as an RGB triplet, a hexadecimal color code, a color name, or a short name.
An RGB triplet is a three-element row vector whose elements specify the intensities of the red, green, and blue components of the color. The intensities must be in the range
[0,1]; for example,[0.4 0.6 0.7].A hexadecimal color code is a character vector or a string scalar that starts with a hash symbol (
#) followed by three or six hexadecimal digits, which can range from0toF. The values are not case sensitive. Thus, the color codes"#FF8800","#ff8800","#F80", and"#f80"are equivalent.
| Color Name | Short Name | RGB Triplet | Hexadecimal Color Code | Appearance |
|---|---|---|---|---|
"red" | "r" | [1 0 0] | "#FF0000" |
|
"green" | "g" | [0 1 0] | "#00FF00" |
|
"blue" | "b" | [0 0 1] | "#0000FF" |
|
"cyan"
| "c" | [0 1 1] | "#00FFFF" |
|
"magenta" | "m" | [1 0 1] | "#FF00FF" |
|
"yellow" | "y" | [1 1 0] | "#FFFF00" |
|
"black" | "k" | [0 0 0] | "#000000" |
|
"white" | "w" | [1 1 1] | "#FFFFFF" |
|
"none" | Not applicable | Not applicable | Not applicable | No color |
This table lists the default color palettes for plots in the light and dark themes.
| Palette | Palette Colors |
|---|---|
Before R2025a: Most plots use these colors by default. |
|
|
|
You can get the RGB triplets and hexadecimal color codes for these palettes using the orderedcolors and rgb2hex functions. For example, get the RGB triplets for the "gem" palette and convert them to hexadecimal color codes.
RGB = orderedcolors("gem");
H = rgb2hex(RGB);Before R2023b: Get the RGB triplets using RGB =
get(groot,"FactoryAxesColorOrder").
Before R2024a: Get the hexadecimal color codes using H =
compose("#%02X%02X%02X",round(RGB*255)).
Example: FaceColor="red"
Example: FaceColor = [0 0.5 0.5]
Example: FaceColor = "#EDB120"
Data Types: double | single | string | char
Line color, specified as an RGB triplet, a hexadecimal color code, a
color name, or a short name. Color is not supported
when plottype is
"histogram".
For a custom color, specify an RGB triplet or a hexadecimal color code.
An RGB triplet is a three-element row vector whose elements specify the intensities of the red, green, and blue components of the color. The intensities must be in the range
[0,1], for example,[0.4 0.6 0.7].A hexadecimal color code is a string scalar or character vector that starts with a hash symbol (
#) followed by three or six hexadecimal digits, which can range from0toF. The values are not case sensitive. Therefore, the color codes"#FF8800","#ff8800","#F80", and"#f80"are equivalent.
Alternatively, you can specify some common colors by name. This table lists the named color options, the equivalent RGB triplets, and the hexadecimal color codes.
| Color Name | Short Name | RGB Triplet | Hexadecimal Color Code | Appearance |
|---|---|---|---|---|
"red" | "r" | [1 0 0] | "#FF0000" |
|
"green" | "g" | [0 1 0] | "#00FF00" |
|
"blue" | "b" | [0 0 1] | "#0000FF" |
|
"cyan"
| "c" | [0 1 1] | "#00FFFF" |
|
"magenta" | "m" | [1 0 1] | "#FF00FF" |
|
"yellow" | "y" | [1 1 0] | "#FFFF00" |
|
"black" | "k" | [0 0 0] | "#000000" |
|
"white" | "w" | [1 1 1] | "#FFFFFF" |
|
"none" | Not applicable | Not applicable | Not applicable | No color |
This table lists the default color palettes for plots in the light and dark themes.
| Palette | Palette Colors |
|---|---|
Before R2025a: Most plots use these colors by default. |
|
|
|
You can get the RGB triplets and hexadecimal color codes for these palettes using the orderedcolors and rgb2hex functions. For example, get the RGB triplets for the "gem" palette and convert them to hexadecimal color codes.
RGB = orderedcolors("gem");
H = rgb2hex(RGB);Before R2023b: Get the RGB triplets using RGB =
get(groot,"FactoryAxesColorOrder").
Before R2024a: Get the hexadecimal color codes using H =
compose("#%02X%02X%02X",round(RGB*255)).
Example: Color="blue"
Example: Color=[0 0 1]
Example: Color="#0000FF"
Data Types: double | single | string | char
Line style, specified as one of the options listed in this table.
| Line Style | Description | Resulting Line |
|---|---|---|
"-" | Solid line |
|
"--" | Dashed line |
|
":" | Dotted line |
|
"-." | Dash-dotted line |
|
"none" | No line | No line |
Example: LineStyle="--"
Data Types: string | char
Line style, specified as one of the options listed in this table.
| Line Style | Description | Resulting Line |
|---|---|---|
"-" | Solid line |
|
"--" | Dashed line |
|
":" | Dotted line |
|
"-." | Dash-dotted line |
|
"none" | No line | No line |
Line width, specified as a positive value in points, where 1 point = 1/72 of an inch. If the line has markers, then the line width also affects the marker edges.
The line width cannot be thinner than the width of a pixel. If you set the line width to a value that is less than the width of a pixel on your system, the line is displayed as one pixel wide.
Example: LineWidth=5
Data Types: double | single
Marker symbol, specified as one of the values listed in this table.
| Marker | Description | Resulting Marker |
|---|---|---|
"o" | Circle |
|
"+" | Plus sign |
|
"*" | Asterisk |
|
"." | Point |
|
"x" | Cross |
|
"_" | Horizontal line |
|
"|" | Vertical line |
|
"square" | Square |
|
"diamond" | Diamond |
|
"^" | Upward-pointing triangle |
|
"v" | Downward-pointing triangle |
|
">" | Right-pointing triangle |
|
"<" | Left-pointing triangle |
|
"pentagram" | Pentagram |
|
"hexagram" | Hexagram |
|
Example: Marker="o"
Data Types: string | char
Indices of data points at which to display markers, specified as a vector of positive integers, or a scalar positive integer. If you do not specify the indices, then MATLAB® displays a marker at every data point.
You cannot specify MarkerIndices when
plottype is
"histogram".
Example: plotResiduals(lme,"probability",MarkerIndices=[1 5
10]) displays a marker at the first, fifth, and tenth data
points in a probability plot.
Example: plotResiduals(lme,"fitted",MarkerIndices=1:3:length(y))
displays a marker every three data points in a plot of the residuals
versus fitted values.
Example: plotResiduals(lme,"lagged",MarkerIndices=5)
displays a marker at the fifth data point in a plot of residuals versus
lagged residuals.
Data Types: double | single
Marker outline color, specified as "auto", an RGB
triplet, a hexadecimal color code, a color name, or a short name. The
default value of "auto" uses the same color as
Color.
For a custom color, specify an RGB triplet or a hexadecimal color code.
An RGB triplet is a three-element row vector whose elements specify the intensities of the red, green, and blue components of the color. The intensities must be in the range
[0,1], for example,[0.4 0.6 0.7].A hexadecimal color code is a string scalar or character vector that starts with a hash symbol (
#) followed by three or six hexadecimal digits, which can range from0toF. The values are not case sensitive. Therefore, the color codes"#FF8800","#ff8800","#F80", and"#f80"are equivalent.
Alternatively, you can specify some common colors by name. This table lists the named color options, the equivalent RGB triplets, and the hexadecimal color codes.
| Color Name | Short Name | RGB Triplet | Hexadecimal Color Code | Appearance |
|---|---|---|---|---|
"red" | "r" | [1 0 0] | "#FF0000" |
|
"green" | "g" | [0 1 0] | "#00FF00" |
|
"blue" | "b" | [0 0 1] | "#0000FF" |
|
"cyan"
| "c" | [0 1 1] | "#00FFFF" |
|
"magenta" | "m" | [1 0 1] | "#FF00FF" |
|
"yellow" | "y" | [1 1 0] | "#FFFF00" |
|
"black" | "k" | [0 0 0] | "#000000" |
|
"white" | "w" | [1 1 1] | "#FFFFFF" |
|
"none" | Not applicable | Not applicable | Not applicable | No color |
This table lists the default color palettes for plots in the light and dark themes.
| Palette | Palette Colors |
|---|---|
Before R2025a: Most plots use these colors by default. |
|
|
|
You can get the RGB triplets and hexadecimal color codes for these palettes using the orderedcolors and rgb2hex functions. For example, get the RGB triplets for the "gem" palette and convert them to hexadecimal color codes.
RGB = orderedcolors("gem");
H = rgb2hex(RGB);Before R2023b: Get the RGB triplets using RGB =
get(groot,"FactoryAxesColorOrder").
Before R2024a: Get the hexadecimal color codes using H =
compose("#%02X%02X%02X",round(RGB*255)).
Example: MarkerEdgeColor="m"
Data Types: double | single | string | char
Marker fill color, specified as "none", an RGB
triplet, a hexadecimal color code, a color name, or a short name.
For a custom color, specify an RGB triplet or a hexadecimal color code.
An RGB triplet is a three-element row vector whose elements specify the intensities of the red, green, and blue components of the color. The intensities must be in the range
[0,1], for example,[0.4 0.6 0.7].A hexadecimal color code is a string scalar or character vector that starts with a hash symbol (
#) followed by three or six hexadecimal digits, which can range from0toF. The values are not case sensitive. Therefore, the color codes"#FF8800","#ff8800","#F80", and"#f80"are equivalent.
Alternatively, you can specify some common colors by name. This table lists the named color options, the equivalent RGB triplets, and the hexadecimal color codes.
| Color Name | Short Name | RGB Triplet | Hexadecimal Color Code | Appearance |
|---|---|---|---|---|
"red" | "r" | [1 0 0] | "#FF0000" |
|
"green" | "g" | [0 1 0] | "#00FF00" |
|
"blue" | "b" | [0 0 1] | "#0000FF" |
|
"cyan"
| "c" | [0 1 1] | "#00FFFF" |
|
"magenta" | "m" | [1 0 1] | "#FF00FF" |
|
"yellow" | "y" | [1 1 0] | "#FFFF00" |
|
"black" | "k" | [0 0 0] | "#000000" |
|
"white" | "w" | [1 1 1] | "#FFFFFF" |
|
"none" | Not applicable | Not applicable | Not applicable | No color |
This table lists the default color palettes for plots in the light and dark themes.
| Palette | Palette Colors |
|---|---|
Before R2025a: Most plots use these colors by default. |
|
|
|
You can get the RGB triplets and hexadecimal color codes for these palettes using the orderedcolors and rgb2hex functions. For example, get the RGB triplets for the "gem" palette and convert them to hexadecimal color codes.
RGB = orderedcolors("gem");
H = rgb2hex(RGB);Before R2023b: Get the RGB triplets using RGB =
get(groot,"FactoryAxesColorOrder").
Before R2024a: Get the hexadecimal color codes using H =
compose("#%02X%02X%02X",round(RGB*255)).
Example: MarkerFaceColor="g"
Data Types: double | single | string | char
Marker size, specified as a positive value in points, where 1 point = 1/72 of an inch.
Example: MarkerSize=9
Data Types: double | single
Output Arguments
Handle to the residual plot, returned as a handle.
Version History
Introduced in R2013bYou can now plot observed versus fitted values by specifying the
plottype input argument as
"observed".
Specify the target axes for the plot by using the ax input
argument.
See Also
MATLAB Command
You clicked a link that corresponds to this MATLAB command:
Run the command by entering it in the MATLAB Command Window. Web browsers do not support MATLAB commands.
웹사이트 선택
번역된 콘텐츠를 보고 지역별 이벤트와 혜택을 살펴보려면 웹사이트를 선택하십시오. 현재 계신 지역에 따라 다음 웹사이트를 권장합니다:
또한 다음 목록에서 웹사이트를 선택하실 수도 있습니다.
사이트 성능 최적화 방법
최고의 사이트 성능을 위해 중국 사이트(중국어 또는 영어)를 선택하십시오. 현재 계신 지역에서는 다른 국가의 MathWorks 사이트 방문이 최적화되지 않았습니다.
미주
- América Latina (Español)
- Canada (English)
- United States (English)
유럽
- Belgium (English)
- Denmark (English)
- Deutschland (Deutsch)
- España (Español)
- Finland (English)
- France (Français)
- Ireland (English)
- Italia (Italiano)
- Luxembourg (English)
- Netherlands (English)
- Norway (English)
- Österreich (Deutsch)
- Portugal (English)
- Sweden (English)
- Switzerland
- United Kingdom (English)