이 번역 페이지는 최신 내용을 담고 있지 않습니다. 최신 내용을 영문으로 보려면 여기를 클릭하십시오.
유사성 기반 잔여 수명 추정
이 예제에서는 전처리를 수행하고, 추세를 도출할 수 있는 특징을 선택하고, 센서 융합으로 건전성 지표를 생성하고, 유사성 RUL 추정기를 훈련시키고, 예지진단 성능을 검증하는 단계를 포함하여 완전한 잔여 수명(RUL) 추정 워크플로를 구축하는 방법을 보여줍니다. 이 예제에서는 PHM2008 challenge dataset의 훈련 데이터를 사용합니다[1].
데이터 준비
데이터셋이 작으므로 전체 성능 저하 데이터를 메모리에 불러올 수 있습니다. 데이터 세트를 현재 디렉터리로 다운로드하고 압축을 풉니다. helperLoadData 헬퍼 함수를 사용하여 훈련 텍스트 파일을 불러오고 타임테이블로 구성된 셀형 배열로 변환합니다. 훈련 데이터는 260개의 RTF(run-to-failure) 시뮬레이션을 포함합니다. 이 측정값 그룹을 "앙상블"이라고 합니다.
url = 'https://ssd.mathworks.com/supportfiles/nnet/data/TurbofanEngineDegradationSimulationData.zip'; websave('TurbofanEngineDegradationSimulationData.zip',url); unzip('TurbofanEngineDegradationSimulationData.zip') degradationData = helperLoadData('train_FD002.txt'); degradationData(1:5)
ans=5×1 cell array
{149×26 table}
{269×26 table}
{206×26 table}
{235×26 table}
{154×26 table}
각 앙상블 멤버는 26개 열을 포함하는 테이블입니다. 열에는 기계 ID, 타임스탬프, 3개의 작동 상태, 21개의 센서 측정값 데이터가 들어 있습니다.
head(degradationData{1}) id time op_setting_1 op_setting_2 op_setting_3 sensor_1 sensor_2 sensor_3 sensor_4 sensor_5 sensor_6 sensor_7 sensor_8 sensor_9 sensor_10 sensor_11 sensor_12 sensor_13 sensor_14 sensor_15 sensor_16 sensor_17 sensor_18 sensor_19 sensor_20 sensor_21
__ ____ ____________ ____________ ____________ ________ ________ ________ ________ ________ ________ ________ ________ ________ _________ _________ _________ _________ _________ _________ _________ _________ _________ _________ _________ _________
1 1 34.998 0.84 100 449.44 555.32 1358.6 1137.2 5.48 8 194.64 2222.7 8341.9 1.02 42.02 183.06 2387.7 8048.6 9.3461 0.02 334 2223 100 14.73 8.8071
1 2 41.998 0.8408 100 445 549.9 1353.2 1125.8 3.91 5.71 138.51 2211.6 8304 1.02 42.2 130.42 2387.7 8072.3 9.3774 0.02 330 2212 100 10.41 6.2665
1 3 24.999 0.6218 60 462.54 537.31 1256.8 1047.5 7.05 9.02 175.71 1915.1 8001.4 0.94 36.69 164.22 2028 7864.9 10.894 0.02 309 1915 84.93 14.08 8.6723
1 4 42.008 0.8416 100 445 549.51 1354 1126.4 3.91 5.71 138.46 2211.6 8304 1.02 41.96 130.72 2387.6 8068.7 9.3528 0.02 329 2212 100 10.59 6.4701
1 5 25 0.6203 60 462.54 537.07 1257.7 1047.9 7.05 9.03 175.05 1915.1 7993.2 0.94 36.89 164.31 2028 7861.2 10.896 0.02 309 1915 84.93 14.13 8.5286
1 6 25.005 0.6205 60 462.54 537.02 1266.4 1048.7 7.05 9.03 175.17 1915.2 7996.1 0.94 36.78 164.27 2028 7868.9 10.891 0.02 306 1915 84.93 14.28 8.559
1 7 42.004 0.8409 100 445 549.74 1347.5 1127.2 3.91 5.71 138.71 2211.6 8307.8 1.02 42.19 130.49 2387.7 8075.5 9.3753 0.02 330 2212 100 10.62 6.4227
1 8 20.002 0.7002 100 491.19 607.44 1481.7 1252.4 9.35 13.65 334.41 2323.9 8709.1 1.08 44.27 315.11 2388 8049.3 9.2369 0.02 365 2324 100 24.33 14.799
뒤에서 수행할 성능 평가를 위해 성능 저하 데이터를 훈련 데이터 세트와 검증 데이터 세트로 분할합니다.
rng('default') % To make sure the results are repeatable numEnsemble = length(degradationData); numFold = 5; cv = cvpartition(numEnsemble, 'KFold', numFold); trainData = degradationData(training(cv, 1)); validationData = degradationData(test(cv, 1));
관심 변수 그룹을 지정합니다.
varNames = string(degradationData{1}.Properties.VariableNames);
timeVariable = varNames{2};
conditionVariables = varNames(3:5);
dataVariables = varNames(6:26);앙상블 데이터의 샘플을 시각화합니다.
nsample = 10;
figure
helperPlotEnsemble(trainData, timeVariable, ...
[conditionVariables(1:2) dataVariables(1:2)], nsample)
작업 영역 군집화
이전 섹션에서 볼 수 있듯이 각 RTF(run-to-failure) 측정값에는 성능 저하 과정을 보여주는 분명한 추세가 없습니다. 이 섹션과 다음 섹션에서는 작동 상태를 사용하여 센서 신호로부터 더 명확한 성능 저하 추세를 추출합니다.
각 앙상블 멤버는 3개의 작동 상태 "op_setting_1", "op_setting_2", "op_setting_3"을 포함합니다. 먼저 각 셀에서 테이블을 추출하여 이들을 하나의 테이블로 결합해 보겠습니다.
trainDataUnwrap = vertcat(trainData{:});
opConditionUnwrap = trainDataUnwrap(:, cellstr(conditionVariables));3차원 산점도 플롯에서 모든 동작점을 시각화합니다. 6개의 영역이 명확히 표시되고 각 영역에서 점이 서로 매우 가까이에 있습니다.
figure helperPlotClusters(opConditionUnwrap)

군집화 기법을 사용하여 6개의 군집을 자동으로 찾아보겠습니다. 여기서는 K-평균 알고리즘을 사용합니다. K-평균은 가장 널리 사용되는 군집화 알고리즘 중 하나이지만, 이 알고리즘을 사용하면 국소 최적값이 발생할 수 있습니다. 서로 다른 초기 상태로 K-평균 군집화 알고리즘을 여러 번 반복한 다음 가장 비용이 낮은 결과를 고르는 것이 좋습니다. 여기서는 알고리즘이 5번 실행되는데, 결과는 동일합니다.
opts = statset('Display', 'final'); [clusterIndex, centers] = kmeans(table2array(opConditionUnwrap), 6, ... 'Distance', 'sqeuclidean', 'Replicates', 5, 'Options', opts);
Replicate 1, 1 iterations, total sum of distances = 0.331378. Replicate 2, 1 iterations, total sum of distances = 0.331378. Replicate 3, 1 iterations, total sum of distances = 0.331378. Replicate 4, 1 iterations, total sum of distances = 0.331378. Replicate 5, 1 iterations, total sum of distances = 0.331378. Best total sum of distances = 0.331378
군집화 결과와 식별된 군집 중심을 시각화합니다.
figure helperPlotClusters(opConditionUnwrap, clusterIndex, centers)
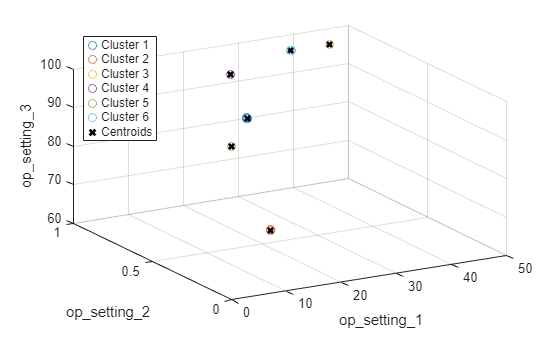
플롯에서 볼 수 있듯이 군집화 알고리즘은 6개의 작업 영역을 성공적으로 찾아냅니다.
작업 영역 정규화
서로 다른 작업 영역으로 그룹화된 측정값을 정규화해 보겠습니다. 먼저, 이전 섹션에서 식별된 작업 영역으로 그룹화된 각 센서 측정값의 평균과 표준편차를 계산합니다.
centerstats = struct('Mean', table(), 'SD', table()); for v = dataVariables centerstats.Mean.(char(v)) = splitapply(@mean, trainDataUnwrap.(char(v)), clusterIndex); centerstats.SD.(char(v)) = splitapply(@std, trainDataUnwrap.(char(v)), clusterIndex); end centerstats.Mean
ans=6×21 table
sensor_1 sensor_2 sensor_3 sensor_4 sensor_5 sensor_6 sensor_7 sensor_8 sensor_9 sensor_10 sensor_11 sensor_12 sensor_13 sensor_14 sensor_15 sensor_16 sensor_17 sensor_18 sensor_19 sensor_20 sensor_21
________ ________ ________ ________ ________ ________ ________ ________ ________ _________ _________ _________ _________ _________ _________ _________ _________ _________ _________ _________ _________
489.05 604.91 1502 1311.2 10.52 15.493 394.33 2319 8785.8 1.26 45.487 371.45 2388.2 8135.5 8.6645 0.03 369.69 2319 100 28.527 17.115
462.54 536.86 1262.7 1050.2 7.05 9.0277 175.43 1915.4 8015.6 0.93992 36.798 164.57 2028.3 7878.6 10.913 0.02 307.34 1915 84.93 14.263 8.5587
445 549.7 1354.4 1127.7 3.91 5.7158 138.63 2212 8328.1 1.0202 42.146 130.55 2388.1 8089.7 9.3745 0.02 331.06 2212 100 10.588 6.3516
491.19 607.56 1485.5 1253 9.35 13.657 334.5 2324 8730 1.0777 44.446 314.88 2388.2 8066.1 9.2325 0.022112 365.37 2324 100 24.455 14.674
518.67 642.67 1590.3 1408.7 14.62 21.61 553.37 2388.1 9063.3 1.3 47.538 521.41 2388.1 8142 8.4419 0.03 393.18 2388 100 38.82 23.292
449.44 555.8 1366.7 1131.5 5.48 8.0003 194.45 2223 8356.4 1.0204 41.981 183.03 2388.1 8072.5 9.3319 0.02 334.22 2223 100 14.831 8.8983
centerstats.SD
ans=6×21 table
sensor_1 sensor_2 sensor_3 sensor_4 sensor_5 sensor_6 sensor_7 sensor_8 sensor_9 sensor_10 sensor_11 sensor_12 sensor_13 sensor_14 sensor_15 sensor_16 sensor_17 sensor_18 sensor_19 sensor_20 sensor_21
__________ ________ ________ ________ __________ _________ ________ ________ ________ __________ _________ _________ _________ _________ _________ __________ _________ _________ __________ _________ _________
1.0346e-11 0.47582 5.8057 8.5359 3.2687e-13 0.0047002 0.67361 0.095286 18.819 0.00017552 0.2538 0.53621 0.098213 16.507 0.038392 6.6965e-16 1.49 0 0 0.14441 0.08598
4.6615e-12 0.36083 5.285 6.8365 1.137e-13 0.0042209 0.44862 0.27006 14.479 0.00087595 0.20843 0.34493 0.28592 13.427 0.043297 3.4003e-16 1.2991 0 3.5246e-12 0.11218 0.066392
0 0.43282 5.6422 7.6564 1.1014e-13 0.0049312 0.43972 0.30478 18.325 0.0015563 0.23965 0.34011 0.3286 16.773 0.037476 1.0652e-15 1.4114 0 0 0.10832 0.064709
1.8362e-11 0.46339 5.7888 7.7768 2.5049e-13 0.0047449 0.6046 0.12828 17.58 0.004193 0.23826 0.49843 0.13156 15.457 0.038886 0.004082 1.4729 0 0 0.13531 0.079592
1.2279e-11 0.4967 6.0349 8.9734 6.573e-14 0.0013408 0.86985 0.07082 19.889 2.7314e-14 0.26508 0.73713 0.070545 17.105 0.037289 6.6619e-16 1.5332 0 0 0.17871 0.1065
1.4098e-11 0.4401 5.6013 7.4461 2.2828e-13 0.0017508 0.47564 0.28634 17.446 0.0019595 0.23078 0.37631 0.30766 15.825 0.038139 3.3656e-16 1.4133 0 0 0.11347 0.067192
각 영역의 통계량은 훈련 데이터를 정규화하는 데 사용할 수 있습니다. 각 앙상블 멤버에 대해 각 행의 동작점을 추출하고 동작점에서 각 군집의 중심까지의 거리를 계산한 다음 가장 가까운 군집 중심을 찾습니다. 그런 다음 각 센서 측정값에서 평균을 뺀 값을 해당 군집의 표준편차로 나눕니다. 거의 일정한 센서 측정값은 잔여 수명 추정에 유용하지 않으므로 표준편차가 0에 가깝다면 정규화된 센서 측정값을 0으로 설정합니다. regimeNormalization 함수에 대한 자세한 내용은 "헬퍼 함수" 섹션을 참조하십시오.
trainDataNormalized = cellfun(@(data) regimeNormalization(data, centers, centerstats), ... trainData, 'UniformOutput', false);
작업 영역별로 정규화된 데이터를 시각화합니다. 정규화 후 일부 센서 측정값의 성능 저하 추세가 드러났습니다.
figure helperPlotEnsemble(trainDataNormalized, timeVariable, dataVariables(1:4), nsample)

추세 특성 분석
이번에는 추세를 가장 잘 도출할 수 있는 센서 측정값을 선택하여 예측을 위한 건전성 지표를 생성합니다. 각 센서 측정값에 대해 선형 성능 저하 모델이 추정되고 신호의 기울기가 평가됩니다.
numSensors = length(dataVariables); signalSlope = zeros(numSensors, 1); warn = warning('off'); for ct = 1:numSensors tmp = cellfun(@(tbl) tbl(:, cellstr(dataVariables(ct))), trainDataNormalized, 'UniformOutput', false); mdl = linearDegradationModel(); % create model fit(mdl, tmp); % train mode signalSlope(ct) = mdl.Theta; end warning(warn);
신호 기울기를 정렬하고 가장 큰 기울기를 갖는 8개의 센서를 선택합니다.
[~, idx] = sort(abs(signalSlope), 'descend');
sensorTrended = sort(idx(1:8))sensorTrended = 8×1
2
3
4
7
11
12
15
17
선택한 추세 도출 가능 센서 측정값을 시각화합니다.
figure helperPlotEnsemble(trainDataNormalized, timeVariable, dataVariables(sensorTrended(3:6)), nsample)
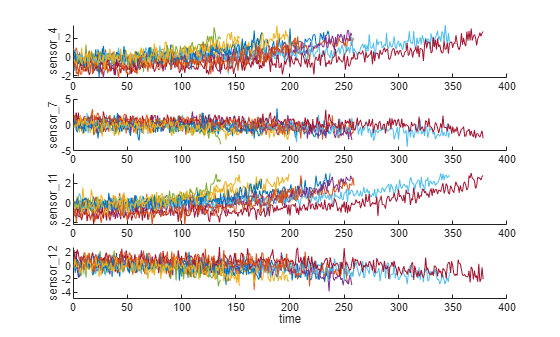
추세를 가장 잘 도출할 수 있는 신호 중 일부는 양의 추세를 보이는 반면, 일부 신호는 음의 추세를 보이는 것을 알 수 있습니다.
건전성 지표 생성하기
이 섹션에서는 센서 측정값을 단일 건전성 지표로 결합하고 이를 사용하여 유사성 모델을 훈련시키는 방법을 살펴봅니다.
모든 RTF(run-to-failure) 데이터는 정상 상태에서 시작한다고 가정합니다. 시작 시점의 건전성 상태에는 값 1이 할당되고 고장 시점의 건전성 상태에는 값 0이 할당됩니다. 건전성 상태는 시간에 따라 1에서 0으로 선형적으로 저하된다고 가정합니다. 이 선형 성능 저하는 센서 값을 융합하는 데 도움이 됩니다. 더욱 정교한 센서 융합 기법은 참고문헌 [2-5]에 설명되어 있습니다.
for j=1:numel(trainDataNormalized) data = trainDataNormalized{j}; rul = max(data.time)-data.time; data.health_condition = rul / max(rul); trainDataNormalized{j} = data; end
건전성 상태를 시각화합니다.
figure
helperPlotEnsemble(trainDataNormalized, timeVariable, "health_condition", nsample)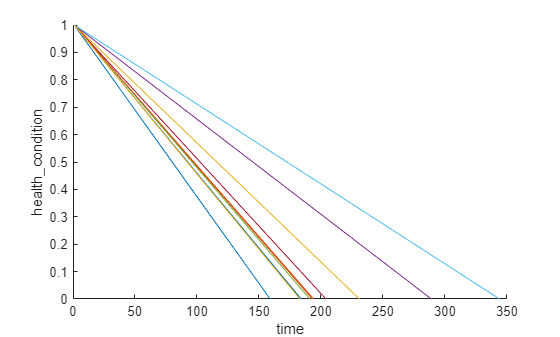
모든 앙상블 멤버의 건전성 상태는 일정하지 않은 성능 저하 속도로 1에서 0까지 변화합니다.
가장 추세가 큰 센서 측정값을 회귀 변수로 사용하여 건전성 상태의 선형 회귀 모델을 피팅합니다.
건전성 상태 ~ 1 + Sensor2 + Sensor3 + Sensor4 + Sensor7 + Sensor11 + Sensor12 + Sensor15 + Sensor17
trainDataNormalizedUnwrap = vertcat(trainDataNormalized{:});
sensorToFuse = dataVariables(sensorTrended);
X = trainDataNormalizedUnwrap{:, cellstr(sensorToFuse)};
y = trainDataNormalizedUnwrap.health_condition;
regModel = fitlm(X,y);
bias = regModel.Coefficients.Estimate(1)bias = 0.5000
weights = regModel.Coefficients.Estimate(2:end)
weights = 8×1
-0.0323
-0.0300
-0.0527
0.0057
-0.0646
0.0054
-0.0431
-0.0379
센서 측정값에 해당 가중치를 곱하여 단일 건전성 지표를 생성합니다.
trainDataFused = cellfun(@(data) degradationSensorFusion(data, sensorToFuse, weights), trainDataNormalized, ... 'UniformOutput', false);
훈련 데이터의 융합된 건전성 지표를 시각화합니다.
figure helperPlotEnsemble(trainDataFused, [], 1, nsample) xlabel('Time') ylabel('Health Indicator') title('Training Data')

여러 센서의 데이터가 단일 건전성 지표로 융합되었습니다. 건전성 지표는 이동평균 필터를 사용하여 평활화됩니다. 자세한 내용은 이전 섹션에서 헬퍼 함수 "degradationSensorFusion"을 참조하십시오.
검증 데이터에 동일한 연산 적용하기
검증 데이터 세트에 대해 영역 정규화 및 센서 융합 과정을 반복합니다.
validationDataNormalized = cellfun(@(data) regimeNormalization(data, centers, centerstats), ... validationData, 'UniformOutput', false); validationDataFused = cellfun(@(data) degradationSensorFusion(data, sensorToFuse, weights), ... validationDataNormalized, 'UniformOutput', false);
검증 데이터의 건전성 지표를 시각화합니다.
figure helperPlotEnsemble(validationDataFused, [], 1, nsample) xlabel('Time') ylabel('Health Indicator') title('Validation Data')
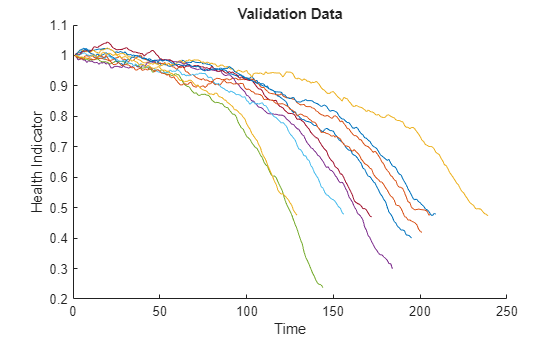
유사성 RUL 모델 구축하기
이번에는 훈련 데이터를 사용하여 잔차 기반 유사성 RUL 모델을 구축합니다. 이 설정에서 모델은 융합된 각 데이터를 2차 다항식을 사용하여 피팅하려고 시도합니다.
데이터 와 데이터 사이의 거리는 잔차의 1-노름을 사용하여 계산됩니다.
여기서 는 기계 의 건전성 지표이고, 는 기계 에서 식별된 2차 다항식 모델을 사용하여 추정된 기계 의 건전성 지표입니다.
유사성 점수는 다음 수식을 사용하여 계산됩니다.
모델은 검증 데이터 세트에서 하나의 앙상블 멤버가 주어지면 훈련 데이터 세트에서 가장 가까운 50개의 앙상블 멤버를 찾고, 50개의 앙상블 멤버를 기반으로 확률 분포를 피팅하고, 분포의 중앙값을 RUL의 추정값으로 사용합니다.
mdl = residualSimilarityModel(... 'Method', 'poly2',... 'Distance', 'absolute',... 'NumNearestNeighbors', 50,... 'Standardize', 1); fit(mdl, trainDataFused);
성능 평가
유사성 RUL 모델을 평가하기 위해 샘플 검증 데이터의 50%, 70% 90%를 사용하여 RUL을 예측합니다.
breakpoint = [0.5, 0.7, 0.9];
validationDataTmp = validationDataFused{3}; % use one validation data for illustration첫 번째 구획점 앞에 있는 검증 데이터를 사용합니다. 이는 수명의 50%에 해당합니다.
bpidx = 1; validationDataTmp50 = validationDataTmp(1:ceil(end*breakpoint(bpidx)),:); trueRUL = length(validationDataTmp) - length(validationDataTmp50); [estRUL, ciRUL, pdfRUL] = predictRUL(mdl, validationDataTmp50);
50%에서 잘린 검증 데이터와 그 최근접이웃을 시각화합니다.
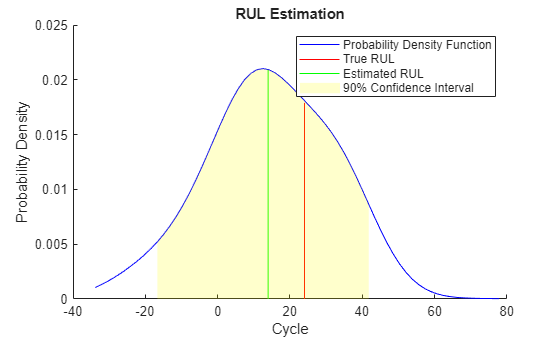
figure compare(mdl, validationDataTmp50);

추정된 RUL과 실제 RUL을 시각화하여 비교하고, 추정된 RUL의 확률 분포를 시각화합니다.
figure helperPlotRULDistribution(trueRUL, estRUL, pdfRUL, ciRUL)

기계가 중간 건전성 단계에 있을 때는 추정된 RUL과 실제 RUL 사이에 상대적으로 큰 오차가 존재합니다. 이 예제에서는 가장 비슷한 10개의 곡선이 시작 지점에 가까이 있지만 고장 상태에 다가가면 두 갈래로 나뉘어서 RUL 분포가 대략적으로 두 개의 모드가 됩니다.
두 번째 구획점 앞에 있는 검증 데이터를 사용합니다. 이는 수명의 70%에 해당합니다.
bpidx = 2; validationDataTmp70 = validationDataTmp(1:ceil(end*breakpoint(bpidx)), :); trueRUL = length(validationDataTmp) - length(validationDataTmp70); [estRUL,ciRUL,pdfRUL] = predictRUL(mdl, validationDataTmp70); figure compare(mdl, validationDataTmp70);
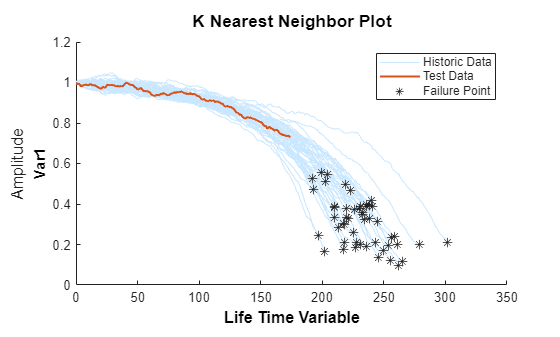
figure helperPlotRULDistribution(trueRUL, estRUL, pdfRUL, ciRUL)
더 많은 데이터가 관측되면 RUL 추정이 개선됩니다.
세 번째 구획점 앞에 있는 검증 데이터를 사용합니다. 이는 수명의 90%에 해당합니다.
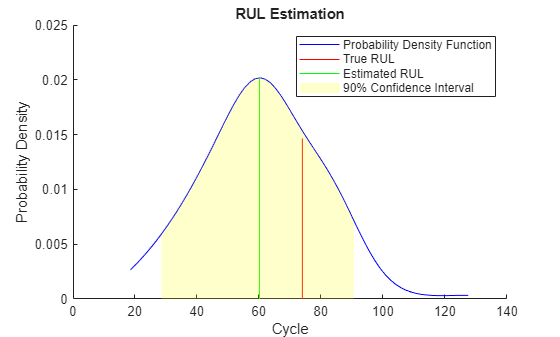
bpidx = 3; validationDataTmp90 = validationDataTmp(1:ceil(end*breakpoint(bpidx)), :); trueRUL = length(validationDataTmp) - length(validationDataTmp90); [estRUL,ciRUL,pdfRUL] = predictRUL(mdl, validationDataTmp90); figure compare(mdl, validationDataTmp90);

figure helperPlotRULDistribution(trueRUL, estRUL, pdfRUL, ciRUL)
이 예제에서는 기계가 고장에 가까워지면 RUL 추정이 한층 더 개선됩니다.
이번에는 전체 검증 데이터 세트에 대해 동일한 평가 절차를 반복하고 각 구획점에 대해 추정된 RUL과 실제 RUL 사이의 오차를 계산합니다.
numValidation = length(validationDataFused); numBreakpoint = length(breakpoint); error = zeros(numValidation, numBreakpoint); for dataIdx = 1:numValidation tmpData = validationDataFused{dataIdx}; for bpidx = 1:numBreakpoint tmpDataTest = tmpData(1:ceil(end*breakpoint(bpidx)), :); trueRUL = length(tmpData) - length(tmpDataTest); [estRUL, ~, ~] = predictRUL(mdl, tmpDataTest); error(dataIdx, bpidx) = estRUL - trueRUL; end end
각 구획점에 대한 오차의 히스토그램을 확률 분포와 함께 시각화합니다.
[pdf50, x50] = ksdensity(error(:, 1)); [pdf70, x70] = ksdensity(error(:, 2)); [pdf90, x90] = ksdensity(error(:, 3)); figure ax(1) = subplot(3,1,1); hold on histogram(error(:, 1), 'BinWidth', 5, 'Normalization', 'pdf') plot(x50, pdf50) hold off xlabel('Prediction Error') title('RUL Prediction Error using first 50% of each validation ensemble member') ax(2) = subplot(3,1,2); hold on histogram(error(:, 2), 'BinWidth', 5, 'Normalization', 'pdf') plot(x70, pdf70) hold off xlabel('Prediction Error') title('RUL Prediction Error using first 70% of each validation ensemble member') ax(3) = subplot(3,1,3); hold on histogram(error(:, 3), 'BinWidth', 5, 'Normalization', 'pdf') plot(x90, pdf90) hold off xlabel('Prediction Error') title('RUL Prediction Error using first 90% of each validation ensemble member') linkaxes(ax)

상자 플롯에 예측 오차를 플로팅하여 중앙값, 25-75 분위수 및 이상값을 시각화합니다.
figure boxplot(error, 'Labels', {'50%', '70%', '90%'}) ylabel('Prediction Error') title('Prediction error using different percentages of each validation ensemble member')

예측 오차의 평균과 표준편차를 계산하고 시각화합니다.
errorMean = mean(error)
errorMean = 1×3
-1.1217 9.5186 9.6540
errorMedian = median(error)
errorMedian = 1×3
1.3798 11.8172 10.3457
errorSD = std(error)
errorSD = 1×3
21.7315 13.5194 12.3083
figure errorbar([50 70 90], errorMean, errorSD, '-o', 'MarkerEdgeColor','r') xlim([40, 100]) xlabel('Percentage of validation data used for RUL prediction') ylabel('Prediction Error') legend('Mean Prediction Error with 1 Standard Deviation Error bar', 'Location', 'south')
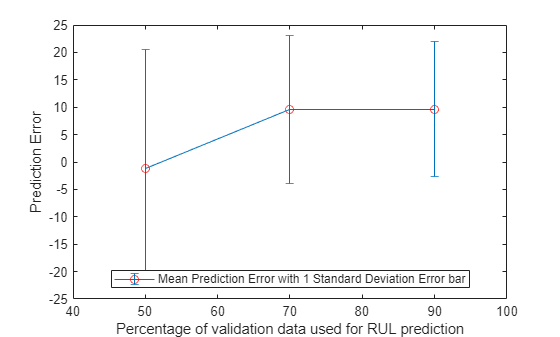
더 많은 데이터가 관측됨에 따라 오차가 0 주변에 더 집중되는 것을(더 적은 이상값) 볼 수 있습니다.
참고 문헌
[1] A. Saxena and K. Goebel (2008). "PHM08 Challenge Data Set", NASA Ames Prognostics Data Repository (http://ti.arc.nasa.gov/project/prognostic-data-repository), NASA Ames Research Center, Moffett Field, CA
[2] Roemer, Michael J., Gregory J. Kacprzynski, and Michael H. Schoeller. "Improved diagnostic and prognostic assessments using health management information fusion." AUTOTESTCON Proceedings, 2001. IEEE Systems Readiness Technology Conference. IEEE, 2001.
[3] Goebel, Kai, and Piero Bonissone. "Prognostic information fusion for constant load systems." Information Fusion, 2005 8th International Conference on. Vol. 2. IEEE, 2005.
[4] Wang, Peng, and David W. Coit. "Reliability prediction based on degradation modeling for systems with multiple degradation measures." Reliability and Maintainability, 2004 Annual Symposium-RAMS. IEEE, 2004.
[5] Jardine, Andrew KS, Daming Lin, and Dragan Banjevic. "A review on machinery diagnostics and prognostics implementing condition-based maintenance." Mechanical systems and signal processing 20.7 (2006): 1483-1510.
헬퍼 함수
function data = regimeNormalization(data, centers, centerstats) % Perform normalization for each observation (row) of the data % according to the cluster the observation belongs to. conditionIdx = 3:5; dataIdx = 6:26; % Perform row-wise operation data{:, dataIdx} = table2array(... rowfun(@(row) localNormalize(row, conditionIdx, dataIdx, centers, centerstats), ... data, 'SeparateInputs', false)); end function rowNormalized = localNormalize(row, conditionIdx, dataIdx, centers, centerstats) % Normalization operation for each row. % Get the operating points and sensor measurements ops = row(1, conditionIdx); sensor = row(1, dataIdx); % Find which cluster center is the closest dist = sum((centers - ops).^2, 2); [~, idx] = min(dist); % Normalize the sensor measurements by the mean and standard deviation of the cluster. % Reassign NaN and Inf to 0. rowNormalized = (sensor - centerstats.Mean{idx, :}) ./ centerstats.SD{idx, :}; rowNormalized(isnan(rowNormalized) | isinf(rowNormalized)) = 0; end function dataFused = degradationSensorFusion(data, sensorToFuse, weights) % Combine measurements from different sensors according % to the weights, smooth the fused data and offset the data % so that all the data start from 1 % Fuse the data according to weights dataToFuse = data{:, cellstr(sensorToFuse)}; dataFusedRaw = dataToFuse*weights; % Smooth the fused data with moving mean stepBackward = 10; stepForward = 10; dataFused = movmean(dataFusedRaw, [stepBackward stepForward]); % Offset the data to 1 dataFused = dataFused + 1 - dataFused(1); end